

The study of sexual selection goes as far as Darwin himself. Since then, elaborate theories concerning both intra- and inter-sexual sexual have been developed, and elegant experiments have been designed to test this body of theory. It may thus come as a surprise that the community is still debating on the correct way to measure simple components of sexual selection, such as the Bateman gradient (i.e., the covariance between the number of matings and the number of offspring)[1,2], or to quantify complex behaviours such as mate choice (the non-random choice of individuals with particular characters as mates)[3,4] and their consequences.
One difficulty in the study of sexual selection is evaluating the consequences of non-random mating. Indeed, when non-random mating is observed in a population, it is often difficult to establish whether such mating pattern leads to i) sexual selection per se (selection pressures favouring certain phenotypes), and/or ii) the non-random association of parental genes in their offspring or not. These two processes differ. In particular, assortative (and disassortative) mating can shape genetic covariances without leading to changes in gene frequencies in the population. Their distinction matters because these two processes lead to different evolutionary outcomes, which can have large ripple effects in the evolution of sexual behaviours, sexual ornamentation, and speciation.
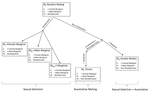
In his paper, entitled “Multi-model inference of non-random mating from an information theoretic approach” [5], Carvajal-Rodríguez tackled this issue. The author generated a simple model in which the consequences of non-random mating can be inferred from information on the population frequencies before and after mating. The procedure is as follows: from the initial population frequencies of phenotypes (or genotypes) of both sexes, the model generates predictions on the frequencies after mating, assuming that particular mating patterns have occurred. This leads to different predictions for the phenotypic (or genotypic) frequencies after mating. The particular mating pattern leading to the best fit with the real frequencies is then identified via a model selection procedure (performing model averaging to combine different mating patterns is also possible).
This study builds on a framework introduced by Carvajal-Rodríguez’s colleagues [6] and encompasses later methodological developments involving the author himself [7]. Compared to early work, the new method proposed by the author builds on the relationship between mating pattern and information [8] to distinguish among scenarios that would lead to non-random mating due to different underlying processes, using simple model selection criterion such as the AICc.
The great asset of the proposed method is that it can be applied to the study of natural populations in which the study of mate choice and sexual selection is notoriously difficult. In the manuscript, the procedure is tested on a population of marine gastropods (Littorina saxatilis). This allows the reader to grasp how the method can be applied to a real system. In fact, anyone can try out the method thanks to the freely available software InfoMating programmed by the author. One important assumption underlying the current method is that the frequencies of unmated individuals do not change during the mating season. If this is not the case, the reader may refer to another publication of the same author which relaxes this assumption [9]. These papers are both instrumental for empiricists interested in testing sexual selection theory.
References
[1] Bateman, A. J. (1948). Intra-sexual selection in Drosophila. Heredity, 2(3), 349-368. doi: 10.1038/hdy.1948.21
[2] Jones, A. G. (2009). On the opportunity for sexual selection, the Bateman gradient and the maximum intensity of sexual selection. Evolution: International Journal of Organic Evolution, 63(7), 1673-1684. doi: 10.1111/j.1558-5646.2009.00664.x
[3] Andersson, M., & Simmons, L. W. (2006). Sexual selection and mate choice. Trends in ecology & evolution, 21(6), 296-302. doi: 10.1016/j.tree.2006.03.015
[4] Kuijper, B., Pen, I., & Weissing, F. J. (2012). A guide to sexual selection theory. Annual Review of Ecology, Evolution, and Systematics, 43, 287-311. doi: 10.1146/annurev-ecolsys-110411-160245
[5] Carvajal-Rodríguez, A. (2019). Multi-model inference of non-random mating from an information theoretic approach. bioRxiv, 305730, ver. 5 peer-reviewed and recommended by PCI Evolutionary Biology. doi: 10.1101/305730
[6] Rolán‐Alvarez, E., & Caballero, A. (2000). Estimating sexual selection and sexual isolation effects from mating frequencies. Evolution, 54(1), 30-36. doi: 10.1111/j.0014-3820.2000.tb00004.x
[7] Carvajal-Rodríguez, A., & Rolan-Alvarez, E. (2006). JMATING: a software for the analysis of sexual selection and sexual isolation effects from mating frequency data. BMC Evolutionary Biology, 6(1), 40. doi: 10.1186/1471-2148-6-40
[8] Carvajal-Rodríguez, A. (2018). Non-random mating and information theory. Theoretical population biology, 120, 103-113. doi: 10.1016/j.tpb.2018.01.003
[9] Carvajal-Rodríguez, A. (2019). A generalization of the informational view of non-random mating: Models with variable population frequencies. Theoretical population biology, 125, 67-74. doi: 10.1016/j.tpb.2018.12.004
DOI or URL of the preprint: 10.1101/305730
Version of the preprint: 3
Dear Sara Magalhaes, I have resubmitted a revised version of the preprint https://doi.org/10.1101/305730 "Multi-model inference of non-random mating from an information theoretic approach".
Thank you very much for your effort. I really appreciate your positive feedback. Please find attached my complete response to your questions/comments. The questions appear in black bold and the answer in normal font. I also attached a pdf with the changes in red.
Dear Antonio,
We have received the revised version of your manuscript, which represents a great improvement relative to the first version, especially concerning the clarity of the message. In line with this, I will not send it our to the reviewers again, but I’d like to ask you to consider still doing a few changes, which I think will make the article even clearer. While you do these changes, I will prepare a recommendation.
Here it goes:
- Line 38: there is still disagreement on its actual definition.
- Line 39: has been challenged.
- Lines 43-44: which ‘various aspects’ are you talking about?
- -Line 44: I would replace “to make things worse” by “Moreover”.
- I would remove all the text that goes from lines 45 to 55. Lines 45-49 repeat a bit what was written before and the paragraph on patterns and processes I think is not needed.
- I would also remove the paragraph from lines 70 to 77, it does not provide much new information.
- Line 82: remove “same”.
- Line 98: the comma should come after “frequencies”, not after “that”.
- Line 101: remove “when measured with matings”.
- I would also remove the paragraph from lines 103-108, as it is quite clear what mating at random is, we don’t need this to be explained.
- Lines 120-123: isn’t it more the opposite: formulating random mating as the zero information model allows expressing the patterns obtained in the other models, right?
- Line 170: remove “the” before “model”.
- Line 210: you haven’t explained what an “information index” is. Please do this before you use the term.
- Line 349: kinds.
- Line 355: remove “as” before “caused”.
- Line 373: replace “be produced” by “occur”, then end the sentence, and state “In fact, in this case…”.
- Line 374: replace “as” by “it is”.
- Line 375: I would put “see also….” Inside the brackets.
- Line 438: another, not other.
- Line 459: replace “at” by “in”.
- Line 466: to apply information criteria to select.
- Line 539: replace “that” by “as”.
- Line 568: to generate.
- Line 627: replace “were” by “was”.
- Line 655: to estimate.
- Lines 659-661: I still think there is insufficient detail in the description of data collection. Do you sample all individuals present? What is a ‘copulation pair’? With which criteria do you distinguish between species? Are these criteria also what you think is involved in sexual selection?
- Lines 715-722: I miss a summary of what happened in the absence of replacement and in the empirical example.
- Lines 723-724: mating tables are not a set-up, please rephrase.
- Line 741: population sizes.
Additional requirements of the managing board:
As indicated in the 'How does it work?’ section and in the code of conduct, please make sure that:
-Data are available to readers, either in the text or through an open data repository such as Zenodo (free), Dryad (to pay) or some other institutional repository. Data must be reusable, thus metadata or accompanying text must carefully describe the data.
-Details on quantitative analyses (e.g., data treatment and statistical scripts in R, bioinformatic pipeline scripts, etc.) and details concerning simulations (scripts, codes) are available to readers in the text, as appendices, or through an open data repository, such as Zenodo, Dryad or some other institutional repository. The scripts or codes must be carefully described so that they can be reused.
-Details on experimental procedures are available to readers in the text or as appendices.
-Authors have no financial conflict of interest relating to the article. The article must contain a "Conflict of interest disclosure" paragraph before the reference section containing this sentence: "The authors of this preprint declare that they have no financial conflict of interest with the content of this article." If appropriate, this disclosure may be completed by a sentence indicating that some of the authors are PCI recommenders: “XXX is one of the PCI XXX recommenders.”
DOI or URL of the preprint: https://doi.org/10.1101/305730
Version of the preprint: 2
Dear Sara Magalhaes, I have resubmitted a revised version of the preprint https://doi.org/10.1101/305730 "Multi-model inference of non-random mating from an information theoretic approach".
Below I am attaching a pdf with my full response to yours and reviewers comments. I fully acknowledge the positive feedback from you and the reviewers. All of the comments has been addressed. Please, find below the detailed answer to the questions. The questions appear in black bold and the answer in normal font. I also attached a pdf with the changes in red.
Dear Author, We have now received three reviews of your manuscript entitled “Multi-model inference of non-random mating from an information theoretic approach”. All reviewers found merit in the approach you’re proposing, but they also raised several issues. I concur with their appreciation and comments. I think this model could be useful to people working on sexual selection. However, I think that the clarity of the manuscript could be improved. In addition to the referees comments, I have a few of my own. I hope that addressing all of them will significantly improve the clearness of your manuscript.
Minor comments:
- Line 11: “to perform”;
- Line 13: please state “in the marine gastropod” before the species name.
- Line 22: explain what you mean by “both kind of patterns”.
- Line 22: remove “models”.
- Line 51: “a posteriori” from what?
- Lines 102-103: replace by “Let a sample have n’ matings”.
- Line 123: replace by “are either known or they need to be estimated”.
- Line 131: “it is convenient”.
- Line 135: remove “Let”.
- Lines 157-158: either you explain which conditions you are referring to or remove this and state it later.
- Line 160: Replace “Following” by “Next”.
- Line 165: “within all others (it is…”.
- Line 177: remove the first “model”.
- Line 186: “if some males have a different value than the other matings”.
- Line 191: “relaxing the first”.
- Line 193: “produce an assortative mating pattern”.
- Line 194: “involves mate choice, which”.
- Line 202: “models”.
- Line 205: “there should be no”.
- Line 215: “all mate types mate at an equal rate”.
- Line 228: “there can be as much”.
- Line 246: “all femate types mate at an equal rate”.
- Line 267-270: I found this section pretty unclear, can you reformulate?
- Line 286: remove “Let”.
- Line 312: “produces”.
- Line 377: “to distinguish”.
- Line 423-424: It would be nice to add a few sentences to explain what you’ll be doing in this section.
- Line 424: “applied to describe”.
- Line 425: “to perform”.
- Line 461: “this indicates”.
- Line 489: the average of what?
- Line 510: “because of”.
- Line 566: please explain “likewise size-assortative mating…”.
- Line 581: “possibly”.
- Line 619: “from these models”.
- Line 623: “SU males do not discriminate between female ecotypes”.
- Line 783: “consists in building”.
I have reviewed the preprint entitle “Multi-model inference of non-random mating from an information theoretic approach” by Antonio Carvajal-Rodriguez (doi: https://doi.org/10.1101/305730). Based on previous work (Carvajal-Rodríguez 2018. Non-random mating and information theory. Theor. Pop. Biol. 120:103-113) the author derived procedures for performing multimodel inference behind from a mating table.
My first comment is that the manuscript is not easy reading and the notation is not always introduced in the right place. For instance, on page 7 m_ij refers to the normalized mating propensity, but its meaning is not clear until next page. I know this was defined in the previous paper, but it would be helpful to have this clear from the beginning.
Starting from the most reduced random mating model, a subset of models are obtained by relaxing some conditions. Mate choice results when the assumption of multiplicability is relaxed. On the other hand, when multiplicability is assume one obtains a pattern of sexual selection. The author then considers several models of increasing complexity and provides the MLE estimates of the different parameters.
Model selection is based on information theory, which was previously shown by the author to provide a valuable framework to make inferences. Simulations suggest that the framework is adequate to estimate the best model, and the procedures were applied to a real case with the gastropod Littorina saxatilis.
Overall, I think the author has done a nice job and his framework can help to understand the role played by the different parameters in a particular case. My main complaint is that the paper is not easy to follow and could be regrettably ignored by some experimentalists. For example, the software MateSim is not user-friendly and it would be very helpful to implement an easier (e.g. windows-based) version. One thing is to perform numerical simulations to explore the parameter space or to test the validity of a given framework, and quite another is to offer a software to be of general use. I suggest the author to put some effort on this last point.
Specific comments
Line 37, page 3: “Mate choice (or intersexual choice) is a process driven by different preference between different mating types”. Not only because the pattern of mating which arises is in part due to mating preferences. Other processes (e.g. spatial distribution of types) can also affect the mate choice.
https://doi.org/10.24072/pci.evolbiol.100186.rev11This manuscript provides a statistical method for estimating which processes (mate choice vs intrasexual competition) underlie patterns of non-random mating, in the case where there is a finite number of discrete phenotypes in each sex. It applies maximum likelihood and model selection methods to the mating table (i.e. the table showing which male-female pairs mated). The statistical framework is sound as far as I could tell, although I would recommend an expert in model selection be invited as reviewer if this has not been done already. However, I think the interpretation of the statistics diverges from mainstream sexual selection theory (in particular in the use of words like ‘mate choice’, ‘intrasexual competition’ and ‘sexual selection’) and is likely to confuse readers who do not understand the formalism.
The manuscript uses the following definitions:
sexual selection: ‘the a posteriori observed change in gene or phenotype frequencies in mated individuals with respect to population frequencies’
intrasexual selection (paraphrased): some individuals (or classes thereof) have uniformly higher mating success than others, independent of the phenotypes of potential partners
mate choice: ‘a process driven by different preference between different mating types’
assortative mating: ‘the a posteriori deviation from random mating within mated individuals’
The most problematic are the definitions of mate choice and intrasexual selection. Most authors (e.g. the classic monograph on sexual selection: Andersson 1994) use ‘intrasexual selection’ in relation to processes like contest and scramble competition, which involve competition among members of one sex without the active involvement of the other sex. In contrast, ‘intersexual selection’ or ‘mate choice’ are generally used where the other sex actively influences the outcome of competition. Doubtless this distinction is hard to make cleanly in all cases, but the current manuscript uses a fundamentally different conceptual taxonomy. E.g. imagine a scenario where males are widely dispersed and never interact with one another. Females travel from male to male and evaluate their phenotypes, mating with preferred males. If some males are preferred by all types of females, the authors would classify this as ‘intrasexual selection’ rather than ‘mate choice’. In their usage, mate choice only occurs if there is variation in preferences among choosers.
The mating table does not contain the relevant information to distinguish between inter- and intra-sexual selection in their traditional senses, whereas it can distinguish between these processes using the authors’ definitions. I’m agnostic about whether the authors’ distinction is biologically useful. Perhaps some people will find it informative for their system. I would consequently recommend that the manuscript be re-written to make clear exactly what is being estimated and making the deviation from common usage clear (or, even better, coming up with some new terms that better capture the meaning of the authors’ definitions).
The definition of sexual selection is less fundamental to understanding this manuscript. However, I should note that most authors define sexual selection as a type of natural selection that arises via competition for mates or fertilisation opportunities (e.g. Andersson 1994; Shuker 2010). Thus, it is important to understand not only how individuals differ in mating success, but also how such differences translate into variation in individual fitness. Under the authors’ definition, ‘sexual selection’ may not be selection at all.
Lastly, the authors’ verbal definition of assortative mating does not quite match up to their mathematical definition. E.g. if some individuals have uniformly higher mating success than others, they may be overrepresented even among mated individuals, indicating a deviation from random mating even among this subpopulation. But the authors would not consider this assortative mating. The verbal definition can be fixed by referring to ‘matings’ rather than ‘mated individuals’.
If these definitional issues were explained clearly, I think this manuscript would make a useful contribution to the literature.
Minor comments:
Line 32: I don’t think ‘variation in mating preferences’ is necessary, just that there exist mating preferences at all (see above). Also intrasexual competition should be mentioned here already.
Line 37: Maybe something like ‘driven by preferences for some traits over others’. The term ‘mating type’ has an existing meaning (molecular characteristics determining the compatibility of gametes) and in any case I don’t think it's a good term here. Maybe just ‘traits’ or ‘phenotypes’.
Line 47: Is this really a ‘decomposition’?
There is an unusual number of typos and other small errors in this manuscript. I started off noting them as I read, but there were so many that I gave up. I would recommend that the authors get a colleague to read it before sending it out for review. I noticed two typos in the maths:
Line 144: Mismatch between ‘theta’ on left-hand side and ‘t’ on right-hand side
Line 226: ‘for all i'
https://doi.org/10.24072/pci.evolbiol.100186.rev13