
GLÉMIN Sylvain
- ECOBIO UMR 6553, CNRS, Rennes, France
- Genome Evolution, Molecular Evolution, Population Genetics / Genomics, Reproduction and Sex
- recommender
Recommendation: 1
Review: 1
Recommendation: 1
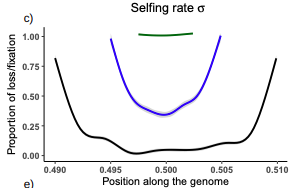
Conditions for maintaining and eroding pseudo-overdominance and its contribution to inbreeding depression
Pseudo-overdominance: how linkage and selection can interact and oppose to purging of deleterious mutations.
Recommended by Sylvain Glémin based on reviews by Yaniv Brandvain, Lei Zhao and 1 anonymous reviewerMost mutations affecting fitness are deleterious and they have many evolutionary consequences. The dynamics and consequences of deleterious mutations are a long-standing question in evolutionary biology and a strong theoretical background has already been developed, for example, to predict the mutation load, inbreeding depression or background selection. One of the classical results is that inbreeding helps purge partially recessive deleterious mutations by exposing them to selection in homozygotes. However, this mainly results from single-locus considerations. When interactions among several, more or less linked, deleterious mutations are taken into account, peculiar dynamics can emerge. One of them, called pseudo-overdominance (POD), corresponds to the maintenance in a population of two (or more) haplotype blocks composed of several recessive deleterious mutations in repulsion that mimics overdominance. Indeed, homozygote individuals for one of the haplotype blocks expose many deleterious mutations to selection whereas they are reciprocally masked in heterozygotes, leading to higher fitness of heterozygotes compared to both homozygotes. A related process, called associative overdominance (AOD) is the effect of such deleterious alleles in repulsion on the linked neutral variation that can be increased by AOD. Although this possibility has been recognized for a long time (Otha and Kimura 1969), it has been mainly considered an anecdotal process. Recently, both theoretical (Zhao and Charlesworth 2016) and genomic analyses (Gilbert et al. 2020) have renewed interest in such a process, suggesting that it could be important in weakly recombining regions of a genome. Donald Waller (2021) - one of the co-authors of the current work - also recently proposed that POD could be quantitatively important with broad implications, and could resolve some unexplained observations such as the maintenance of inbreeding depression in highly selfing species. Yet, a proper theoretical framework analysing the effect of inbreeding on POD was lacking.
In this theoretical work, Diala Abu Awad and Donald Waller (2022) addressed this question through an elegant combination of analytical predictions and intensive multilocus simulations. They determined the conditions under which POD can be maintained and how long it could resist erosion by recombination, which removes the negative association between deleterious alleles (repulsion) at the core of the mechanism. They showed that under tight linkage, POD regions can persist for a long time and generate substantial segregating load and inbreeding depression, even under inbreeding, so opposing (for a while) to the purging effect. They also showed that background selection can affect the genomic structure of POD regions by rapidly erasing weak POD regions but maintaining strong POD regions (i.e with many tightly linked deleterious alleles).
These results have several implications. They can explain the maintenance of inbreeding depression despite inbreeding (as anticipated by Waller 2021), which has implications for the evolution of mating systems. If POD can hardly emerge under high selfing, it can persist from an outcrossing ancestor long after the transition towards a higher selfing rate and could explain the maintenance of mixed mating systems(which is possible with true overdominance, see Uyenoyama and Waller 1991). The results also have implications for genomic analyses, pointing to regions of low or no recombination where POD could be maintained, generating both higher diversity and heterozygosity than expected and variance in fitness. As structural variations are likely widespread in genomes with possible effects on suppressing recombination (Mérot et al. 2020), POD regions should be checked more carefully in genomic analyses (see also Gilbert et al. 2020).
Overall, this work should stimulate new theoretical and empirical studies, especially to assess how quantitatively strong and widespread POD can be. It also stresses the importance of properly considering genetic linkage genome-wide, and so the role of recombination landscapes in determining patterns of diversity and fitness effects.
Awad DA, Waller D (2022) Conditions for maintaining and eroding pseudo-overdominance and its contribution to inbreeding depression. bioRxiv, 2021.12.16.473022, ver. 3 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2021.12.16.473022
Gilbert KJ, Pouyet F, Excoffier L, Peischl S (2020) Transition from Background Selection to Associative Overdominance Promotes Diversity in Regions of Low Recombination. Current Biology, 30, 101-107.e3. https://doi.org/10.1016/j.cub.2019.11.063
Mérot C, Oomen RA, Tigano A, Wellenreuther M (2020) A Roadmap for Understanding the Evolutionary Significance of Structural Genomic Variation. Trends in Ecology & Evolution, 35, 561–572. https://doi.org/10.1016/j.tree.2020.03.002
Ohta T, Kimura M (1969) Linkage disequilibrium at steady state determined by random genetic drift and recurrent mutation. Genetics, 63, 229–238. https://doi.org/10.1093/genetics/63.1.229
Uyenoyama MK, Waller DM (1991) Coevolution of self-fertilization and inbreeding depression II. Symmetric overdominance in viability. Theoretical Population Biology, 40, 47–77. https://doi.org/10.1016/0040-5809(91)90046-I
Waller DM (2021) Addressing Darwin’s dilemma: Can pseudo-overdominance explain persistent inbreeding depression and load? Evolution, 75, 779–793. https://doi.org/10.1111/evo.14189
Zhao L, Charlesworth B (2016) Resolving the Conflict Between Associative Overdominance and Background Selection. Genetics, 203, 1315–1334. https://doi.org/10.1534/genetics.116.188912
Review: 1

Unraveling genetic load dynamics during biological invasion: insights from two invasive insect species
The genetic load of invasive population: how little do we know ?
Recommended by Quentin Rougemont based on reviews by Sylvain Glémin and 2 anonymous reviewersWe live both in a worrying and fascinating time. Worrying because human-induced global change has dramatic consequences on biodiversity around the world. Fascinating because these changes enable us to witness evolutionary processes unfolding on relatively short time scales. One such process is biological invasion. An intriguing evolutionary question is to understand which factor facilitates the success of an invasive species. In particular, serial bottlenecks at the expanding front should reduce the effective population size and decrease genetic diversity. Theoretically, this will increase the fixation of deleterious mutations due to the effect of genetic drift and overall affect the evolutionary potential of the invading species. In the short term, reduced genetic diversity and inbreeding in small populations increases the number of recessive deleterious variants exposed in a homozygous state. This may generate a reduction in mean fitness of the population. However, in the long term and under specific demographic scenarios recessive deleterious alleles may be more efficiently removed by purifying selection. Such purging may explain the success of invasion by reducing inbreeding depression and minimizing loss of fitness. Here, Lombaert et al. estimate the genetic load in two invasive insect species, a predator species, the harlequin ladybird (Harmonia axyridis) and a crop pest species, the western corn rootworm (Diabrotica virgifera virgifera).
The authors smartly took advantage of a pool-seq transcriptome-based exome capture method to estimate genetic load and assess the purge hypothesis using standard population genetic statistics, such as the ratio of nonsynonymous over synonymous expected heterozygosity, the frequency of derived alleles, and their excess or deficit.
The results revealed different patterns in the two species:
In the western corn rootworm, the authors find a clear signal of reduced genetic diversity in invasive populations. This was associated with a slightly reduced genetic load. However, there was only marginal evidence of purging regarding the most deleterious mutations, and in a single population, with moderately deleterious variants being weakly purged, as theoretically expected.
In the harlequin ladybird, in contrast, the reduction of genetic diversity in invasive populations has been small, a result related to the mild severity of the bottlenecks. In this species, the authors found a tendency toward fixation of the genetic load and no signal of purging.
Such results are intriguing, showing that different species seem to exhibit contrasted fate of genetic load. Differences in the invasion history and ecology of the species may explain these patterns. This is one of the first studies to use a population genomics approach to study the genetic load associated with biological invasion. Future studies based on whole genome data collected at the individual level across multiple species are needed to better understand the dynamics of genetic load during biological invasion and to draw more general conclusions. Advances in forward simulations may also be used to shed light on the evolution of the genetic load at different stages of the invasion process and under different strengths of bottlenecks.
References
Eric Lombaert, Aurelie Blin, Barbara Porro, Thomas Guillemaud, Julio S Bernal, Gary Chang, Natalia Kirichenko, Thomas W Sappington, Stefan Toepfer, Emeline Deleury (2025) Unraveling genetic load dynamics during biological invasion: insights from two invasive insect species. bioRxiv, ver.3 peer-reviewed and recommended by PCI Evol Biol https://doi.org/10.1101/2024.09.02.610743