
SCHIFFELS Stephan
- Department for Archaeogenetics, Max Planck Institute for Evolutionary Anthropology, Leipzig, Germany
- Bioinformatics & Computational Biology, Human Evolution, Molecular Evolution, Population Genetics / Genomics
- recommender
Recommendation: 1
Reviews: 0
Recommendation: 1
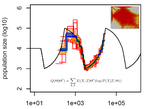
Limits and Convergence properties of the Sequentially Markovian Coalescent
Review and Assessment of Performance of Genomic Inference Methods based on the Sequentially Markovian Coalescent
Recommended by Stephan Schiffels based on reviews by 3 anonymous reviewersThe human genome not only encodes for biological functions and for what makes us human, it also encodes the population history of our ancestors. Changes in past population sizes, for example, affect the distribution of times to the most recent common ancestor (tMRCA) of genomic segments, which in turn can be inferred by sophisticated modelling along the genome.
A key framework for such modelling of local tMRCA tracts along genomes is the Sequentially Markovian Coalescent (SMC) (McVean and Cardin 2005, Marjoram and Wall 2006) . The problem that the SMC solves is that the mosaic of local tMRCAs along the genome is unknown, both in their actual ages and in their positions along the genome. The SMC allows to effectively sum across all possibilities and handle the uncertainty probabilistically. Several important tools for inferring the demographic history of a population have been developed built on top of the SMC, including PSMC (Li and Durbin 2011), diCal (Sheehan et al 2013), MSMC (Schiffels and Durbin 2014), SMC++ (Terhorst et al 2017), eSMC (Sellinger et al. 2020) and others.
In this paper, Sellinger, Abu Awad and Tellier (2020) review these SMC-based methods and provide a coherent simulation design to comparatively assess their strengths and weaknesses in a variety of demographic scenarios (Sellinger, Abu Awad and Tellier 2020). In addition, they used these simulations to test how breaking various key assumptions in SMC methods affects estimates, such as constant recombination rates, or absence of false positive SNP calls.
As a result of this assessment, the authors not only provide practical guidance for researchers who want to use these methods, but also insights into how these methods work. For example, the paper carefully separates sources of error in these methods by observing what they call “Best-case convergence” of each method if the data behaves perfectly and separating that from how the method applies with actual data. This approach provides a deeper insight into the methods than what we could learn from application to genomic data alone.
In the age of genomics, computational tools and their development are key for researchers in this field. All the more important is it to provide the community with overviews, reviews and independent assessments of such tools. This is particularly important as sometimes the development of new methods lacks primary visibility due to relevant testing material being pushed to Supplementary Sections in papers due to space constraints. As SMC-based methods have become so widely used tools in genomics, I think the detailed assessment by Sellinger et al. (2020) is timely and relevant.
In conclusion, I recommend this paper because it bridges from a mere review of the different methods to an in-depth assessment of performance, thereby addressing both beginners in the field who just seek an initial overview, as well as experienced researchers who are interested in theoretical boundaries and assumptions of the different methods.
References
[1] Li, H., and Durbin, R. (2011). Inference of human population history from individual whole-genome sequences. Nature, 475(7357), 493-496. doi: https://doi.org/10.1038/nature10231
[2] Marjoram, P., and Wall, J. D. (2006). Fast"" coalescent"" simulation. BMC genetics, 7(1), 16. doi: https://doi.org/10.1186/1471-2156-7-16
[3] McVean, G. A., and Cardin, N. J. (2005). Approximating the coalescent with recombination. Philosophical Transactions of the Royal Society B: Biological Sciences, 360(1459), 1387-1393. doi: https://doi.org/10.1098/rstb.2005.1673
[4] Schiffels, S., and Durbin, R. (2014). Inferring human population size and separation history from multiple genome sequences. Nature genetics, 46(8), 919-925. doi: https://doi.org/10.1038/ng.3015
[5] Sellinger, T. P. P., Awad, D. A., Moest, M., and Tellier, A. (2020). Inference of past demography, dormancy and self-fertilization rates from whole genome sequence data. PLoS Genetics, 16(4), e1008698. doi: https://doi.org/10.1371/journal.pgen.1008698
[6] Sellinger, T. P. P., Awad, D. A. and Tellier, A. (2020) Limits and Convergence properties of the Sequentially Markovian Coalescent. bioRxiv, 2020.07.23.217091, ver. 3 peer-reviewed and recommended by PCI Evolutionary Biology. doi: https://doi.org/10.1101/2020.07.23.217091
[7] Sheehan, S., Harris, K., and Song, Y. S. (2013). Estimating variable effective population sizes from multiple genomes: a sequentially Markov conditional sampling distribution approach. Genetics, 194(3), 647-662. doi: https://doi.org/10.1534/genetics.112.149096
[8] Terhorst, J., Kamm, J. A., and Song, Y. S. (2017). Robust and scalable inference of population history from hundreds of unphased whole genomes. Nature genetics, 49(2), 303-309. doi: https://doi.org/10.1038/ng.3748