
Making sense of vertebrate sirtuin genes
How many sirtuin genes are out there? evolution of sirtuin genes in vertebrates with a description of a new family member
Abstract
Recommendation: posted 22 September 2022, validated 29 September 2022
Delsuc, F. (2022) Making sense of vertebrate sirtuin genes. Peer Community in Evolutionary Biology, 100151. https://doi.org/10.24072/pci.evolbiol.100151
Recommendation
Sirtuin proteins are class III histone deacetylases that are involved in a variety of fundamental biological functions mostly related to aging. These proteins are located in different subcellular compartments and are associated with different biological functions such as metabolic regulation, stress response, and cell cycle control [1]. In mammals, the sirtuin gene family is composed of seven paralogs (SIRT1-7) grouped into four classes [2]. Due to their involvement in maintaining cell cycle integrity, sirtuins have been studied as a way to understand fundamental mechanisms governing longevity [1]. Indeed, the downregulation of sirtuin genes with aging seems to explain much of the pathophysiology that accumulates with aging [3]. Biomedical studies have thus explored the potential therapeutic implications of sirtuins [4] but whether they can effectively be used as molecular targets for the treatment of human diseases remains to be demonstrated [1]. Despite this biomedical interest and some phylogenetic analyses of sirtuin paralogs mostly conducted in mammals, a comprehensive evolutionary analysis of the sirtuin gene family at the scale of vertebrates was still lacking.
In this preprint, Opazo and collaborators [5] took advantage of the increasing availability of whole-genome sequences for species representing all main groups of vertebrates to unravel the evolution of the sirtuin gene family. To do so, they undertook a phylogenomic approach in its original sense aimed at improving functional predictions by evolutionary analysis [6] in order to inventory the full vertebrate sirtuin gene repertoire and reconstruct its precise duplication history. Harvesting genomic databases, they extracted all predicted sirtuin proteins and performed phylogenetic analyses based on probabilistic inference methods. Maximum likelihood and Bayesian analyses resulted in well-resolved and congruent phylogenetic trees dividing vertebrate sirtuin genes into three major clades. These analyses also revealed an additional eighth paralog that was previously overlooked because of its restricted phyletic distribution. This newly identified sirtuin family member (named SIRT8) was recovered with unambiguous statistical support as a sister-group to the SIRT3 clade. Comparative genomic analyses based on conserved gene synteny confirmed that SIRT8 was present in all sampled non-amniote vertebrate genomes (cartilaginous fish, bony fish, coelacanth, lungfish, and amphibians) except cyclostomes. SIRT8 has thus most likely been lost in the last common ancestor of amniotes (mammals, reptiles, and birds). Discovery of such previously unknown genes in vertebrates is not completely surprising given the plethora of high-quality genomes now available. However, this study highlights the importance of considering a broad taxonomic sampling to infer evolutionary patterns of gene families that have been mostly studied in mammals because of their potential importance for human biology.
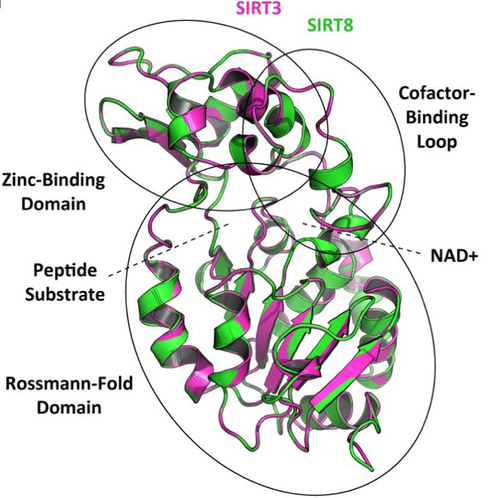
Based on its phylogenetic position as closely related to SIRT3 within class I, it could be predicted that the newly identified SIRT8 paralog likely has a deacetylase activity and is probably located in mitochondria. To test these evolutionary predictions, Opazo and collaborators [5] conducted further bioinformatics analyses and functional experiments using the elephant shark (Callorhinchus milii) as a model species. RNAseq expression data were analyzed to determine tissue-specific transcription of sirtuin genes in vertebrates, including SIRT8 found to be mainly expressed in the ovary, which suggests a potential role in biological processes associated with reproduction. The elephant shark SIRT8 protein sequence was used with other vertebrates for comparative analyses of protein structure modeling and subcellular localization prediction both pointing to a probable mitochondrial localization. The protein localization and its function were further characterized by immunolocalization in transfected cells, and enzymatic and functional assays, which all confirmed the prediction that SIRT8 proteins are targeted to the mitochondria and have deacetylase activity. The extensive experimental efforts made in this study to shed light on the function of this newly discovered gene are both rare and highly commendable.
Overall, this work by Opazo and collaborators [5] provides a comprehensive phylogenomic study of the sirtuin gene family in vertebrates based on detailed evolutionary analyses using state-of-the-art phylogenetic reconstruction methods. It also illustrates the power of adopting an integrative comparative approach supplementing the reconstruction of the duplication history of the gene family with complementary functional experiments in order to elucidate the function of the newly discovered SIRT8 family member. These results provide a reference phylogenetic framework for the evolution of sirtuin genes and the further functional characterization of the eight vertebrate paralogs with potential relevance for understanding the cellular biology of aging and its associated diseases in human.
References
[1] Vassilopoulos A, Fritz KS, Petersen DR, Gius D (2011) The human sirtuin family: Evolutionary divergences and functions. Human Genomics, 5, 485. https://doi.org/10.1186/1479-7364-5-5-485
[2] Yamamoto H, Schoonjans K, Auwerx J (2007) Sirtuin Functions in Health and Disease. Molecular Endocrinology, 21, 1745–1755. https://doi.org/10.1210/me.2007-0079
[3] Morris BJ (2013) Seven sirtuins for seven deadly diseases ofaging. Free Radical Biology and Medicine, 56, 133–171. https://doi.org/10.1016/j.freeradbiomed.2012.10.525
[4] Bordo D Structure and Evolution of Human Sirtuins. Current Drug Targets, 14, 662–665. http://dx.doi.org/10.2174/1389450111314060007
[5] Opazo JC, Vandewege MW, Hoffmann FG, Zavala K, Meléndez C, Luchsinger C, Cavieres VA, Vargas-Chacoff L, Morera FJ, Burgos PV, Tapia-Rojas C, Mardones GA (2022) How many sirtuin genes are out there? evolution of sirtuin genes in vertebrates with a description of a new family member. bioRxiv, 2020.07.17.209510, ver. 5 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2020.07.17.209510
[6] Eisen JA (1998) Phylogenomics: Improving Functional Predictions for Uncharacterized Genes by Evolutionary Analysis. Genome Research, 8, 163–167. https://doi.org/10.1101/gr.8.3.163
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was supported by the Fondo Nacional de Desarrollo Científico y Tecnológico from Chile (FONDECYT 1210471) and The Millennium Nucleus of Ion Channel-Associated Diseases is a Millennium Nucleus of the Millennium Scientific Initiative, National Agency of Research and Development (ANID, NCN19_168), Ministry of Science, Technology, Knowledge, and Innovation, Chile to J.C.O., Fondo Nacional de Desarrollo Científico y Tecnológico from Chile (FONDECYT 1211481) to G.A.M., National Science Foundation (EPS-0903787, DBI-1262901, and DEB-1354147) to F.G.H., Financiamiento Basal No. ANID/BASAL/ACE210009 to P.V.B. and No. ANID/BASAL/FB210008 to P.V.B. and C.T.R., Fondap-Ideal Grant No. 15150003 and ANID-Millennium Science Initiative Program-Center ICM-ANID ICN2021_002 to L.V.C., Fondo Nacional de Desarrollo Científico y Tecnológico from Chile (FONDECYT 1180957) to F.J.M., Vicerrectoría de Investigación, Desarrollo y Creación Artística, Universidad Austral de Chile to J.C.O., F.J.M., L.V.C., G.A.M
Reviewed by Nicolas Leurs, 24 Aug 2022
The authors have answered to my comments either by modifying the text and/or figures accordingly, by justifying their choices with convincing arguments or by providing additional information or analyses that support their conclusions. Therefore I am favorable to the publication of this article as is, and congratulate the authors on their work.
https://doi.org/10.24072/pci.evolbiol.100151.rev21Reviewed by anonymous reviewer 1, 20 Aug 2022
I am happy to recommend that this manuscript be accepted.
https://doi.org/10.24072/pci.evolbiol.100151.rev22Reviewed by Filipe Castro, 19 Aug 2022
Dear Frédéric,
The authors have modified the manuscript accordingly. I have nothing further. A very elegant study indeed.
Best wishes
Filipe
https://doi.org/10.24072/pci.evolbiol.100151.rev23Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2020.07.17.209510
Version of the preprint: 3
Author's Reply, 12 Aug 2022
Decision by Frédéric Delsuc, posted 07 Jul 2022
Dear Drs. Juan Opazo and Gonzalo Mardones,
I now have received three reviews from colleagues that have carefuly read your preprint entitled "How many sirtuin genes are out there? Evolution of sirtuin genes in vertebrates with a description of a new family member" submitted for evaluation by Peer Community in Evolutionary Biology.
As you will see in their detailed review reports, all three reviewers found the manuscript interesting and clearly written. I agree with them that your study provides a comprehensive study of this gene family based on detailed evolutionary analyses and I share their appreciation of the efforts you deployed to shed light on the potential function of the newly discovered sirtuin SIRT3-like paralog through functional experiments. All three reviewers also provide valuable suggestions that I think will improve the clarity of the study.
The following points are particularly worth addressing in your revision:
1. More details are needed regarding the construction of the alignment. In particular, the length of the alignment and whether or not it has been processed with a site filtering method before phylogenetic analysis should be indicated. If site filtering has not been performed, as suggested by one of the reviewer, the effect of applying a method such as HMMCleaner on phylogenetic inference might be worth exploring.
2. Two reviewers required additional explanation/clarification regarding the phylogenetic position of the lamprey sequence to the SIRT3 clade and the potential reasons why this sequence is particularly difficult to place in the tree. I agree with them that the potential effect of the compositional bias of the lamprey sequence should be further explored and discussed.
3. Finally, two reviewers had questions concerning the name of the newly identified SIRT3-like paralog and suggest renaming the gene independently of its similarity to SIRT3 by perhaps simply calling it SIRT8. It would be good to hear your informed opinion on this nomenclatural issue.
I am looking forward to read the revised version of the manuscript.
Best regards,
Frédéric Delsuc on the behalf of PCI Evol Biol
Reviewed by Filipe Castro, 23 Jun 2022
Opazo and collaborators provide in this manuscript an exhaustive and comprehensive evolutionary analysis of the SIRT gene family in vertebrates species. The study is indeed elegant, clarifying the origin of SIRT3-like genes, and highlighting the importance of taxonomic sampling to infer evolutionary patterns (incidentally, is the gene present in lungfishes?). The authors further analyze and explore the function of this novel gene utilizing an ample set of approaches – e.g. gene expression, in vivo analysis, modeling and protein function. I have no real comments and congratulate the authors for their work! I understand the authors prevent from speculation but is there a reasonable scenario for both the specific retention and loss of SIRT3-like in these disparate lineages? A minor doubt relates with nomenclature: I’m not an expert but I wonder whether it would be interesting/possible/advisable (?) to rename the gene independently of its similarity to SIRT3?
Reviewed by Nicolas Leurs, 13 Jun 2022
Reviewed by anonymous reviewer 1, 08 Jun 2022
This study rexamines and clarifies the evolutionary history and repertoire of the multifunctional vertebrate Sirtuin gene family. The authors find evidence for a new family member that has been lost in amniotes. Discovery of such previously unknown genes is not especially unusual in vertebrate gene family studies that broadly sample genome-data from across vertebrate phylogeny, however the extensive effort made in this study to shine a light on the function of this new gene is both rare and highly commendable.
I have no major concerns about the study, although I note that I lack sufficient expertise to critique the functional experiments, and only a few minor points that the authors may wish to consider:
1. Regarding the placement of SIRT3 the authors state "This situation could be explained by the sequence itself, instead of the well-known compositional bias of the cyclostome genomes" (line 136-137). Is it not possible that the opposite to this is occuring here, with compositional bias affecting the placement of the lamprey sequence but not the hagfish? Similarly (although it seems unlikely) from the results shown it seems that paralogy of the lamprey sequence to SIRT3 cannot be ruled out?
2. It might be worth noting that the 'high transcription levels' in ovary for the newly discovered SIRT3-like, although still interesting, lack biological replicates at least within species (especially given the Zebrafish findings).
https://doi.org/10.24072/pci.evolbiol.100549.rev13









