
An unusual suspect: the mutation landscape as a determinant of local variation in nucleotide diversity
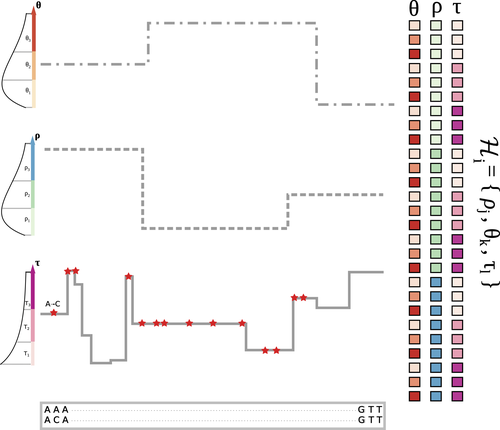
The landscape of nucleotide diversity in Drosophila melanogaster is shaped by mutation rate variation
Abstract
Recommendation: posted 13 April 2023, validated 13 April 2023
Racimo, F. (2023) An unusual suspect: the mutation landscape as a determinant of local variation in nucleotide diversity. Peer Community in Evolutionary Biology, 100636. https://doi.org/10.24072/pci.evolbiol.100636
Recommendation
Sometimes, important factors for explaining biological processes fall through the cracks, and it is only through careful modeling that their importance eventually comes out to light. In this study, Barroso and Dutheil introduce a new method based on the sequentially Markovian coalescent (SMC, Marjoran and Wall 2006) for jointly estimating local recombination and coalescent rates along a genome. Unlike previous SMC-based methods, however, their method can also co-estimate local patterns of variation in mutation rates.
This is a powerful improvement which allows them to tackle questions about the reasons for the extensive variation in nucleotide diversity across the chromosomes of a species - a problem that has plagued the minds of population geneticists for decades (Begun and Aquadro 1992, Andolfatto 2007, McVicker et al., 2009, Pouyet and Gilbert 2021). The authors find that variation in de novo mutation rates appears to be the most important factor in determining nucleotide diversity in Drosophila melanogaster. Though seemingly contradicting previous attempts at addressing this problem (Comeron 2014), they take care to investigate and explain why that might be the case.
Barroso and Dutheil have also taken care to carefully explain the details of their new approach and have carried a very thorough set of analyses comparing competing explanations for patterns of nucleotide variation via causal modeling. The reviewers raised several issues involving choices made by the authors in their analysis of variance partitioning, the proper evaluation of the role of linked selection and the recombination rate estimates emerging from their model. These issues have all been extensively addressed by the authors, and their conclusions seem to remain robust. The study illustrates why the mutation landscape should not be ignored as an important determinant of local variation in genetic diversity, and opens up questions about the generalizability of these results to other organisms.
REFERENCES
Andolfatto, P. (2007). Hitchhiking effects of recurrent beneficial amino acid substitutions in the Drosophila melanogaster genome. Genome research, 17(12), 1755-1762. https://doi.org/10.1101/gr.6691007
Barroso, G. V., & Dutheil, J. Y. (2021). The landscape of nucleotide diversity in Drosophila melanogaster is shaped by mutation rate variation. bioRxiv, 2021.09.16.460667, ver. 3 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2021.09.16.460667
Begun, D. J., & Aquadro, C. F. (1992). Levels of naturally occurring DNA polymorphism correlate with recombination rates in D. melanogaster. Nature, 356(6369), 519-520. https://doi.org/10.1038/356519a0
Comeron, J. M. (2014). Background selection as baseline for nucleotide variation across the Drosophila genome. PLoS Genetics, 10(6), e1004434. https://doi.org/10.1371/journal.pgen.1004434
Marjoram, P., & Wall, J. D. (2006). Fast" coalescent" simulation. BMC genetics, 7, 1-9. https://doi.org/10.1186/1471-2156-7-16
McVicker, G., Gordon, D., Davis, C., & Green, P. (2009). Widespread genomic signatures of natural selection in hominid evolution. PLoS genetics, 5(5), e1000471. https://doi.org/10.1371/journal.pgen.1000471
Pouyet, F., & Gilbert, K. J. (2021). Towards an improved understanding of molecular evolution: the relative roles of selection, drift, and everything in between. Peer Community Journal, 1, e27. https://doi.org/10.24072/pcjournal.16
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was supported by a grant from the German Research Foundation (Deutsche Forschungsgemeinschaft) attributed to JYD, within the priority program (SPP) 1590 “probabilistic structures in evolution”
Evaluation round #2
DOI or URL of the preprint: https://doi.org/10.1101/2021.09.16.460667
Version of the preprint: 2
Author's Reply, 02 Apr 2023
Dear Recommander,
We thank you and the reviewer for pointing out ways in which we could improve the presentation of our results. We were able to incorporate all suggestions into the updated version of our manuscript: Figure S1 now includes the demography inferred by iSMC (piecewise constant within time intervals, mapped from splines parameters) as well as the smoothed sketch used for coalescent simulations; Greek letters are now used in the legends of Figures 4 and 5; Panel titles for Figure 5 have been updated to reflect that block lengths of constant mutation rate represent averages from a geometric distribution; Tables 1 and S2 now include visual guides. Clarifications regarding these changes were added in lines 696-703. Comments from reviewer 1 are addressed below.
We are thankful that the reviewer considers our work a contribution to the field, and we agree that our model has shortcomings. For example, in line 532 we mention how it can be improved in the future. We also reiterate that the binning of genomic landscapes into windows is independent from genome-wide parameter estimation and do not see it as a step back in terms of interpretability (we tried to clarify this further in lines 688-694). On the contrary, that our linear model explains >99% of the distribution of diversity along the genome is evidence that our framework is adequate to describe the effects of drift, mutation, recombination and linked selection on patterns of DNA variation. Thus we can use typical procedures like ANOVA and standardized coefficients to assess the impact of each micro-evolutionary mechanism on levels of diversity. These are rather easy to interpret in terms of relative importance. In the end, our findings are not incompatible with the existing literature because previous studies on linked selection focused on its relative importance after removing the effect of mutation rate variation, and also because there are, in fact, studies highlighting the importance of mutation rate variation, although for the most part they have been wrongfully ignored. We provide citations to some of these throughout the main text.
Decision by Fernando Racimo, posted 20 Mar 2023, validated 20 Mar 2023
Dear authors,
The reviewers have read your replies to them and are (mostly) satisfied with them. Thank you for your answers to the queries by me and the reviewers.
I don't think it will be necessary for the reviewers to see you manuscript again, as long as the points below are addressed.
Best,
Fernando
1. It's unclear whether Supplemental Figure S1 is a direct result of your inference, or a "sketch" (inspired by what?) that you then used in simulations. As reviewer 2 rightly points out (R2.3), there should be a global TMRCA emerging from this analysis, even if not the central focus of your study. If this can't be produced for some reason, a better explanation for that reason should be available in the text.
2. Please improve the resolution of Supplemental Figure S1 so that the x-axis can be read.
3. Figure 5 warrants more explanation as to what is being depicted here exactly. For example, does "mu block ~500kb" imply that the mutation rate was simulated so as to vary in blocks of (approximately?) 500 kb? In the text it says exactly 500 kb. Also, could you replace "TMRCA" for "tau", and use the greek symbols in the figure as you use in the text?
4. Can you draw horizontal lines in Table 1 and Table S2 to help the reader figure out when one model ends and another begins?
5. Can you address this comment by the anonymous reviewer in your text? "Couldn’t lower autocorrelation instead result not from frequent variation in recombination rate window-to-window, but relatively few windows with extreme shifts in recombination rate relative to their neighboring windows?"
Reviewed by David Castellano, 17 Mar 2023
Barroso and Dutheil have addressed my main concerns and clarified the issues I raised in my previous review. I do not have any further comments.
https://doi.org/10.24072/pci.evolbiol.100636.rev21Reviewed by anonymous reviewer 1, 17 Mar 2023
Evaluation round #1
DOI or URL of the preprint: https://doi.org/10.1101/2021.09.16.460667
Version of the preprint: 1
Author's Reply, 16 Feb 2023
Decision by Fernando Racimo, posted 09 Dec 2022, validated 13 Dec 2022
In this manuscript, Barroso & Dutheil present a new method for co-estimating local recombination rates, local mutation rates and local effective population sizes along the genome, and then apply it to a Drosophila melanogaster haploid genome panel from Zambia. They find a strong role for local variation in mutation rate on variation in local patterns of diversity along the genome - a finding that appears to reach contradictory conclusions to previous approaches to the question of the major determinants of local diversity. The paper is well written, and I agree with the reviewers that the approach is innovative and elegant. I also think the methodology is very well explained. I have some concerns about the robustness of the biological conclusions, and their dependence on particular decisions by the authors. The first reviewers' point about the size of analysis windows should be further explored, and the authors could do a more thorough test into the role of linked selection using simulations. The second reviewer also raised some important points about how the chosen shape for the DFE could influence parameters estimation, and about how the recombination rate estimates could be compared to empirical estimates. I would be happy to recommend this manuscript once these concerns are addressed.
Reviewed by anonymous reviewer 1, 06 Dec 2022
Reviewed by David Castellano, 28 Nov 2022
In this manuscript, Barroso & Dutheil propose an extension of a statistical method that jointly infers the genomic landscape of genealogies (or "local Ne"), recombination rates and mutation rates. They benchmark the method with simulations and apply it to Drosophila melanogaster from Zambia. They find that, at the genomic window lengths that they analyze (50Kb, 200Kb and 1Mb), the mutation landscape seems to be the most important determinant of the levels of genetic diversity along the Drosophila genome. This conclusion is somehow contradicting Comeron 2014, where he concluded that the genetic diversity landscape is mostly affected by linked selection (or tau, or TMRCA, using Barroso & Dutheil terminology). However, the authors do a good job of reasoning why both studies seem to reach contradictory conclusions. This manuscript is relevant to the population genomics community because it makes available a powerful tool and it brings back to the spot light the mutation landscape. I agree that the mutation landscape is often an overlooked ingredient to explain the genetic diversity landscape within a genome.
I've divided this revision into 4 sections.
1. Is the science sound, with a logical narrative and well-supported results and conclusions?
The manuscript follows a logical narrative and the methods are sound. The simulations and benchmarking are convincing. The literature context provided in the introduction and discussion is very helpful (below I suggest a couple of more papers to back up some ideas tho). However, some aspects require further clarification.
1.1 I do not think that the partial R^2 is a good way to assess the relative importance of each variable (mutation rate, recombination rate and TMRCA) on the levels of genetic diversity. The standardized regression coefficients, which are the regression coefficients obtained from estimating a model on the standardized variables (mean = 0, standard deviation = 1), are better suited for this job IMO. I would also suggest reporting the variance inflation factors in Table 1.
1.2 I wonder why the authors do not try to validate their mutation rate and recombination rate estimates using empirical measures. I understand that perhaps empirical measures of the mutation rate (using mutation accumulation inbreed lines?) might be hard to find, but the empirical recombination landscape from Comeron is publicly available. Moreover, how the global or genome-wide TMRCA inferred with the new method compares to previous demographic estimates in this population? If they are not similar then what can be the cause?
1.3 Related to the previous point. For future implementations maybe in the Discussion. Could it be possible to plug-in empirical mutation and recombination landscapes to infer the local TMRCA (or "local Ne")?
1.4 Regarding the DFE of Drosophila. The authors simulate a shape = 1 (row 654), this is equivalent to an exponential distribution and will produce way more weakly deleterious mutations than the ones expected when the shape = 0.3-0.4 (which is the value more commonly estimated in the literature for this species, see https://academic.oup.com/mbe/article/35/11/2685/5078937 and others). This excess of weakly deleterious mutations could explain, I believe, the results explained from rows 406 to 415.
1.5 Suggested literature.
1.5.1 In here (https://www.ncbi.nlm.nih.gov/pmc/articles/PMC1575889/) Spencer et al. 2006 use "wavelet techniques to identify correlations acting at different scales" which in essence is very similar to Barroso & Dutheil work.
1.5.2 In here (https://onlinelibrary.wiley.com/doi/10.1002/bies.201200150) Martincorena and Luscombe 2012 review "the main forces driving the evolution of local mutation rates and identify the main limiting factors" which might be relevant for sentences in rows 495-500.
1.5.3 Bear in mind that Castellano et al. 2018a was already published in GBE "long" ago. The cited preprinted version might be outdated.
1.6 Could the authors provide some intuition about the "five mutation rate classes, five recombination rate classes and 30 coalescent time intervals" used? Why this setting and not another one? How relevant is this choice in downstream analyses?
2. Is there enough info to allow verifying and reproducing the data?
The supplementary information, plus the scripts, are easy to access.
3. Are there obscure passages that you (or a potential reader) can’t go through?
3.1 I do not understand the first sentence of the paragraph starting at row 154.
3.2 In figures 4 and 5 it might be helpful to scale the y-axes in log units?
3.3 Typo at row 245 "the contribution of contribution"?
4. Potential extra analysis only if interesting enough to the recommender and/or author:
Relevant to rows 346-355. I am just curious to know how much tau (or TMRCA) varies along the genome in the absence of selection compared to the presence of selection. Could the authors show some density plots in both scenarios? I think it is often assumed that in the absence of selection genetic diversity is entirely explained by the mutation landscape. Still, it seems that the TMRCA (and genetic diversity) can vary along the genome stochastically in the presence of recombination. This is an interesting finding that should be further highlighted.
https://doi.org/10.24072/pci.evolbiol.100636.rev12









