
GANDON Sylvain
- CEFE, CNRS, Montpellier, France
- Evolutionary Ecology, Evolutionary Epidemiology, Evolutionary Theory, Experimental Evolution, Life History, Other, Phenotypic Plasticity
- recommender
Recommendations: 2
Reviews: 0
Recommendations: 2
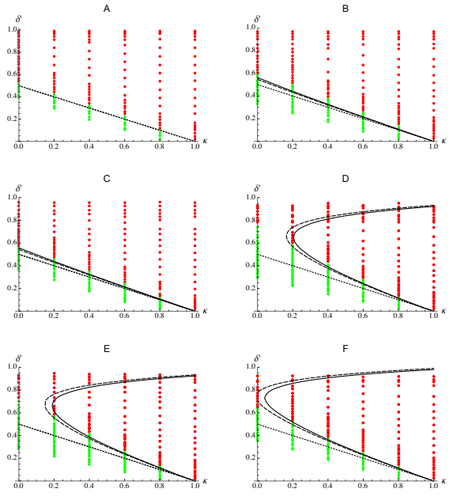
Epistasis, inbreeding depression and the evolution of self-fertilization
Epistasis and the evolution of selfing
Recommended by Sylvain Gandon based on reviews by Nick Barton and 1 anonymous reviewerThe evolution of selfing results from a balance between multiple evolutionary forces. Selfing provides an "automatic advantage" due to the higher efficiency of selfers to transmit their genes via selfed and outcrossed offspring. Selfed offspring, however, may suffer from inbreeding depression. In principle the ultimate evolutionary outcome is easy to predict from the relative magnitude of these two evolutionary forces [1,2]. Yet, several studies explicitly taking into account the genetic architecture of inbreeding depression noted that these predictions are often too restrictive because selfing can evolve in a broader range of conditions [3,4].
The present work by Abu Awad and Roze [5] provides an analytic understanding of these results. Abu Awad and Roze analyse the evolution of selfing in a multilocus model where some loci are coding for selfing while others are under direct selection. The evolution of selfing depends on (i) the classical benefit of selfing (automatic advantage), (ii) the cost of selfing due to inbreeding depression, (iii) the association between the loci coding for selfing and the loci under direct selection (likely to be positive because selfing is expected to be found in better purged genetic backgrounds) and (iv) the association between the loci coding for selfing and the linkage between loci under selection (this final term depends on the magnitude and the type of epistasis). Because these last two terms depend on genetic associations they are expected to play in when selection is strong and recombination is small. These last two terms explain why selfing is evolving under a range of conditions which is broader than predicted by earlier theoretical models. The match between the approximations for the different terms acting on the evolution of selfing and individual based simulations (for different fitness landscapes) is very convincing. In particular, this analysis also yields new results on the effect of different types of epistasis on inbreeding depression.
Another remarkable and important feature of this work is its readability. The analysis of multilocus models rely on several steps and approximations that often result in overwhelmingly complex papers. Abu Awad and Roze’s paper [5] is dense but it provides a very clear and comprehensive presentation of the interplay between multiple evolutionary forces acting on the evolution of selfing.
References
[1] Holsinger, K. E., Feldman, M. W., and Christiansen, F. B. (1984). The evolution of self-fertilization in plants: a population genetic model. The American Naturalist, 124(3), 446-453. doi: 10.1086/284287
[2] Lande, R., and Schemske, D. W. (1985). The evolution of self‐fertilization and inbreeding depression in plants. I. Genetic models. Evolution, 39(1), 24-40. doi: 10.1111/j.1558-5646.1985.tb04077.x
[3] Charlesworth, D., Morgan, M. T., and Charlesworth, B. (1990). Inbreeding depression, genetic load, and the evolution of outcrossing rates in a multilocus system with no linkage. Evolution, 44(6), 1469-1489. doi: 10.1111/j.1558-5646.1990.tb03839.x
[4] Uyenoyama, M. K., and Waller, D. M. (1991). Coevolution of self-fertilization and inbreeding depression I. Mutation-selection balance at one and two loci. Theoretical population biology, 40(1), 14-46. doi: 10.1016/0040-5809(91)90045-H
[5] Abu Awad, D. and Roze, D. (2020). Epistasis, inbreeding depression and the evolution of self-fertilization. bioRxiv, 809814, ver. 4 peer-reviewed and recommended by PCI Evol Biol. doi: 10.1101/809814
Human adaptation of Ebola virus during the West African outbreak
Ebola evolution during the 2013-2016 outbreak
Recommended by Sylvain Gandon and Sébastien LionThe Ebola virus (EBOV) epidemic that started in December 2013 resulted in around 28,000 cases and more than 11,000 deaths. Since the emergence of the disease in Zaire in 1976 the virus had produced a number of outbreaks in Africa but until 2013 the reported numbers of human cases had never risen above 500. Could this exceptional epidemic size be due to the spread of a human-adapted form of the virus?
The large mutation rate of the virus [1-2] may indeed introduce massive amounts of genetic variation upon which selection may act. Several earlier studies based on the accumulation of genome sequences sampled during the epidemic led to contrasting conclusions. A few studies discussed evidence of positive selection on the glycoprotein that may be linked to phenotypic variations on infectivity and/or immune evasion [3-4]. But the heterogeneity in the transmission of some lineages could also be due to environmental heterogeneity and/or stochasticity. Most studies could not rule out the null hypothesis of the absence of positive selection and human adaptation [1-2 and 5].
In a recent experimental study, Urbanowicz et al. [6] chose a different method to tackle this question. A phylogenetic analysis of genome sequences from viruses sampled in West Africa revealed the existence of two main lineages (one with a narrow geographic distribution in Guinea, and the other with a wider geographic distribution) distinguished by a single amino acid substitution in the glycoprotein of the virus (A82V), and of several sub-lineages characterised by additional substitutions. The authors used this phylogenetic data to generate a panel of mutant pseudoviruses and to test their ability to infect human and fruit bat cells. These experiments revealed that specific amino acid substitutions led to higher infectivity of human cells, including A82V. This increased infectivity on human cells was associated with a decreased infectivity in fruit bat cell cultures. Since fruit bats are likely to be the reservoir of the virus, this paper indicates that human adaptation may have led to a specialization of the virus to a new host.
An accompanying paper in the same issue of Cell by Diehl et al. [7] reports results that confirm the trend identified by Urbanowicz et al. [6] and further indicate that the increased infectivity of A82V is specific for primate cells. Diehl et al. [7] also report some evidence for higher virulence of A82V in humans. In other words, the evolution of the virus may have led to higher abilities to infect and to kill its novel host. This work thus confirms the adaptive potential of RNA virus and the ability of Ebola to specialize to a novel host. In this context, the availability of an effective vaccine against the disease is particularly welcome [8].
The study of Urbanowicz et al. [6] is also remarkable because it illustrates the need of experimental approaches for the study of phenotypic variation when inference methods based on phylodynamics fail to extract a clear biological message. The analysis of genomic evolution is still in its infancy and there is a need for new theoretical developments to help detect more rapidly candidate mutations involved in adaptations to new environmental conditions.
References
[1] Gire, S.K., Goba, A., Andersen, K.G., Sealfon, R.S.G., Park, D.J., Kanneh, L., Jalloh, S., Momoh, M., Fullah, M., Dudas, G., et al. (2014). Genomic surveillance elucidates Ebola virus origin and transmission during the 2014 outbreak. Science 345, 1369–1372. doi: 10.1126/science.1259657
[2] Hoenen, T., Safronetz, D., Groseth, A., Wollenberg, K.R., Koita, O.A., Diarra, B., Fall, I.S., Haidara, F.C., Diallo, F., Sanogo, M., et al. (2015). Mutation rate and genotype variation of Ebola virus from Mali case sequences. Science 348, 117–119. doi: 10.1126/science.aaa5646
[3] Liu, S.-Q., Deng, C.-L., Yuan, Z.-M., Rayner, S., and Zhang, B. (2015). Identifying the pattern of molecular evolution for Zaire ebolavirus in the 2014 outbreak in West Africa. Infection, Genetics and Evolution 32, 51–59. doi: 10.1016/j.meegid.2015.02.024
[4] Holmes, E.C., Dudas, G., Rambaut, A., and Andersen, K.G. (2016). The evolution of Ebola virus: Insights from the 2013–2016 epidemic. Nature 538, 193–200. doi: 10.1038/nature19790
[5] Azarian, T., Lo Presti, A., Giovanetti, M., Cella, E., Rife, B., Lai, A., Zehender, G., Ciccozzi, M., and Salemi, M. (2015). Impact of spatial dispersion, evolution, and selection on Ebola Zaire Virus epidemic waves. Scientific Reports. 5, 10170. doi: 10.1038/srep10170
[6] Urbanowicz, R.A., McClure, C.P., Sakuntabhai, A., Sall, A.A., Kobinger, G., Müller, M.A., Holmes, E.C., Rey, F.A., Simon-Loriere, E., and Ball, J.K. (2016). Human adaptation of Ebola virus during the West African outbreak. Cell 167, 1079–1087. doi: 10.1016/j.cell.2016.10.013
[7] Diehl, W.E., Lin, A.E., Grubaugh, N.D., Carvalho, L.M., Kim, K., Kyawe, P.P., McCauley, S.M., Donnard, E., Kucukural, A., McDonel, P., et al. (2016). Ebola virus glycoprotein with increased infectivity dominated the 2013-2016 epidemic. Cell 167, 1088–1098. doi: 10.1016/j.cell.2016.10.014
[8] Henao-Restrepo, A.M., Camacho, A., Longini, I.M., Watson, C.H., Edmunds, W.J., Egger, M., Carroll, M.W., Dean, N.E., Diatta, I., Doumbia, M., et al. (2016). Efficacy and effectiveness of an rVSV-vectored vaccine in preventing Ebola virus disease: final results from the Guinea ring vaccination, open-label, cluster-randomised trial (Ebola Ça Suffit!). The Lancet 389, 505-518. doi: 10.1016/S0140-6736(16)32621-6