
Epistasis and the evolution of selfing
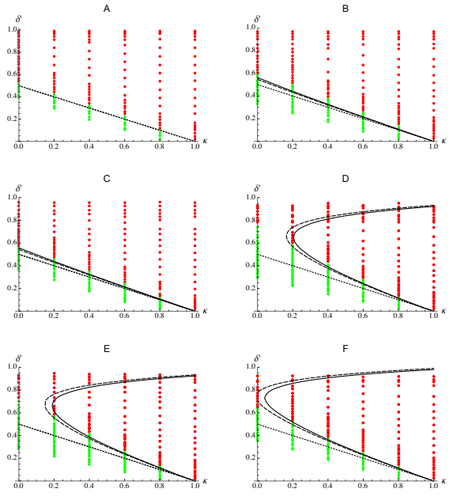
Epistasis, inbreeding depression and the evolution of self-fertilization
Abstract
Recommendation: posted 08 February 2020, validated 17 February 2020
Gandon, S. (2020) Epistasis and the evolution of selfing. Peer Community in Evolutionary Biology, 100093. https://doi.org/10.24072/pci.evolbiol.100093
Recommendation
The evolution of selfing results from a balance between multiple evolutionary forces. Selfing provides an "automatic advantage" due to the higher efficiency of selfers to transmit their genes via selfed and outcrossed offspring. Selfed offspring, however, may suffer from inbreeding depression. In principle the ultimate evolutionary outcome is easy to predict from the relative magnitude of these two evolutionary forces [1,2]. Yet, several studies explicitly taking into account the genetic architecture of inbreeding depression noted that these predictions are often too restrictive because selfing can evolve in a broader range of conditions [3,4].
The present work by Abu Awad and Roze [5] provides an analytic understanding of these results. Abu Awad and Roze analyse the evolution of selfing in a multilocus model where some loci are coding for selfing while others are under direct selection. The evolution of selfing depends on (i) the classical benefit of selfing (automatic advantage), (ii) the cost of selfing due to inbreeding depression, (iii) the association between the loci coding for selfing and the loci under direct selection (likely to be positive because selfing is expected to be found in better purged genetic backgrounds) and (iv) the association between the loci coding for selfing and the linkage between loci under selection (this final term depends on the magnitude and the type of epistasis). Because these last two terms depend on genetic associations they are expected to play in when selection is strong and recombination is small. These last two terms explain why selfing is evolving under a range of conditions which is broader than predicted by earlier theoretical models. The match between the approximations for the different terms acting on the evolution of selfing and individual based simulations (for different fitness landscapes) is very convincing. In particular, this analysis also yields new results on the effect of different types of epistasis on inbreeding depression.
Another remarkable and important feature of this work is its readability. The analysis of multilocus models rely on several steps and approximations that often result in overwhelmingly complex papers. Abu Awad and Roze’s paper [5] is dense but it provides a very clear and comprehensive presentation of the interplay between multiple evolutionary forces acting on the evolution of selfing.
References
[1] Holsinger, K. E., Feldman, M. W., and Christiansen, F. B. (1984). The evolution of self-fertilization in plants: a population genetic model. The American Naturalist, 124(3), 446-453. doi: 10.1086/284287
[2] Lande, R., and Schemske, D. W. (1985). The evolution of self‐fertilization and inbreeding depression in plants. I. Genetic models. Evolution, 39(1), 24-40. doi: 10.1111/j.1558-5646.1985.tb04077.x
[3] Charlesworth, D., Morgan, M. T., and Charlesworth, B. (1990). Inbreeding depression, genetic load, and the evolution of outcrossing rates in a multilocus system with no linkage. Evolution, 44(6), 1469-1489. doi: 10.1111/j.1558-5646.1990.tb03839.x
[4] Uyenoyama, M. K., and Waller, D. M. (1991). Coevolution of self-fertilization and inbreeding depression I. Mutation-selection balance at one and two loci. Theoretical population biology, 40(1), 14-46. doi: 10.1016/0040-5809(91)90045-H
[5] Abu Awad, D. and Roze, D. (2020). Epistasis, inbreeding depression and the evolution of self-fertilization. bioRxiv, 809814, ver. 4 peer-reviewed and recommended by PCI Evol Biol. doi: 10.1101/809814
The recommender in charge of the evaluation of the article and the reviewers declared that they have no conflict of interest (as defined in the code of conduct of PCI) with the authors or with the content of the article. The authors declared that they comply with the PCI rule of having no financial conflicts of interest in relation to the content of the article.
This work was supported by the French Agence Nationale de la Recherche (project SEAD, ANR-13-ADAP0011 and project SexChange, ANR-14-CE02-0001). Diala Abu Awad was partly funded by the TUM University Foundation Fellowship.
Evaluation round #1
DOI or URL of the preprint: https://www.biorxiv.org/content/10.1101/809814v1
Author's Reply, 04 Feb 2020
Dear Editors, we thank the editor and reviewers for very helpful comments which we think have helped to improve our manuscript. We have tried to address all of them as explained in the attached file, and posted a new version of the manuscript on bioRxiv (https://www.biorxiv.org/content/10.1101/809814v2). We hope that the manuscript can be recommended on PCI. Best regards, Denis Roze
Decision by Sylvain Gandon, posted 11 Dec 2019
This is a very impressive piece of work on a technically difficult but biologically very relevant question. The model clarifies the role different components of epistasis on inbreeding depression and on the evolution of selfing. The introduction replaces this model in among previous studies that have addressed this subject.
I am very much willing to recommend this preprint but it would be great to use the comments from both reviewers to improve the manuscript. The paper is beautifully written but very dense. There are two main parts (inbreeding depression and evolution of selfing) but many subcases. I don't know if it is feasible but it would be great to provide a way (an additional figure?) to summarize some aspects of your work to the less theoretically-oriented readers. You may want to try provide a schematic description of some features of your model. Visualizing the different components of epistasis would be useful given their importance in your model. Also it would be great to see what it means for the fitness landscape to assume that epistasis is fixed or distributed. You may want to introduce sooner in the text the deleterious mutation rate (not explicitly mentioned before line 276).
Reviewed by anonymous reviewer 1, 09 Dec 2019
Reviewed by Nick Barton, 30 Nov 2019
This paper analyses selection on modifiers of selfing. It extends previous work by using a very general model of selection which includes arbitrary epistasis. The analysis is complex and technically sophisticated, and yet presented very clearly; for three specific models of epistasis, comparisons with simulation show that the approximations are accurate. The key achievement is to write the change in modifier frequency as the sum of distinct terms, which can be attributed to different processes, and (at least partly) related to components of fitness variance. Thus, Abu Awad and Roze have made an impressive contribution to our understanding of how selfing evolves.
The approach is based on the QLE approximation, and is similar to Barton (1995), who looked at the much simpler problem of evolution of recombination with random mating. My main comment is that a bit more could be done to relate these results to the earlier analysis. Specifically:
- epistatic coefficients are assumed of order eps^2, relative to directional coefficients, which are of order eps. This is necessary if the variance components are to be of the same order, and arises because there are L^2 pairwise coefficients, but only L directional coefficients. This emerges in S1 but is not explained in the main text.
- Deleterious mutations are assumed rare, but this assumption is not made consistently: sometimes, terms like (1-pj) appear, which should be close to 1; conversely, epistatic terms necessarily involve pj*p_k, which would seem to be second order. I think that the analysis is actually OK, but it needs a more careful explanation.
Specific points:
Eq 1 - Would random variation in selfing rate make any difference. I guess not, but it is not immediately obvious.
Eq 7 - Maybe this will come later, but there should be a comment on the consequences of setting up selfing rate as a quantitative trait, rather than looking at invasion of a specific allele. I suspect that the qualitative outcome is the same, but again, this is not obvious. (see line 220 also)
198 - This choice of a generalised function leads to higher-order epistasis, and so will require some approximation.
238 - Write lsigma=10 here
274 "the different terms in (19) contribute multiplicatively" is ambiguous (though I can guess what it might mean)
276 - Worth commenting on why, under these assumptions, only the mean # of bad mutations (nd) matters.
** 279 - One should explain the four terms in (25), which should be possible. The crucial assumption that deleterious alleles are rare deserves more prominence. It is not clear that this assumption is used consistently, since one would expect second order terms to arise from aj and aj,j; the last two terms are necessarily second order. It may be that the n^2 factor makes the latter two p^2 terms more important than the neglected second-order contributions to the first two terms, but that needs attention.
350 - In (33) it seems that p is no longer assumed small. Clarify when that assumption is used, and when not!
- In (36) do i, j still refer to loci for selfing & for fitness, respectively?
- The independence of (46) from Q is surprising. Why is this so?
472 - This statement may reveal a fundamental difficulty. Can the a of different order be assumed to be of the same order? (as it were; cf Barton 1995, and above)?
- Can one interpret the various components in terms of the change in fitness mean and variance due to recombination, etc? (see 648)
664 How does the advantage of outcrossing mediated by Hill-Robertson effects appear here? This involves terms of order aj ak ajk which do not appear here; they may have been neglected because of different assumptions about the order of terms. This is quite confusing.
484 - Here and elsewhere, can one consistently include the small term aj in the denominator, whilst neglecting corrections of this order elsewhere?
636 - cite Otto on the importance of variance in epistasis across pairs of loci. (I think she made tis point in a n Annual Review with Feldman, arguing that it necessarily weakened selection for recombination)
- It may be unwise to rely on colour in the figures - this makes it hard for people to read them offline.
S1 - Is there a numerical check that these selection coefficients are correctly calculated?
S5 - The selfing rate is the average of the two alleles, yet in the text, it is defined as their sum.










