
BRANDVAIN Yaniv
- , University of Minnesota - Twin Citites, St Paul, United States of America
- Evolutionary Theory, Hybridization / Introgression, Population Genetics / Genomics, Quantitative Genetics, Speciation
- recommender
Recommendation: 1
Reviews: 2
Recommendation: 1

Parallel pattern of differentiation at a genomic island shared between clinal and mosaic hybrid zones in a complex of cryptic seahorse lineages
Genomic parallelism in adaptation to orthogonal environments in sea horses
Recommended by Yaniv Brandvain based on reviews by 3 anonymous reviewersStudies in speciation genomics have revealed that gene flow is quite common, and that despite this, species can maintain their distinct environmental adaptations. Although researchers are still elucidating the genomic mechanisms by which species maintain their adaptations in the face of gene flow, this often appears to involve few diverged genomic regions in otherwise largely undifferentiated genomes. In this preprint [1], Riquet and colleagues investigate the genetic structuring and patterns of parallel evolution in the long-snouted seahorse.
Before investigating specific SNPs plausibly associated with adaptation, the authors first describe genome-wide population structure in the long-snouted seahorse. This species is split into five phenotypically similar, but genetically distinct populations. Two populations reside in the Atlantic Ocean and are geographically structured with one north of the Iberian peninsula and the other around the Iberian peninsula. Two other populations are found in the Mediterranean Sea and are structured by the environment as they correspond to marine and lagoon environments. The genetic clustering of lagoon populations in the Mediterranean, despite the substantial geographic distance between them is quite impressive, and worthy of further study. Finally, a fifth population resides in a lagoon-like habitat in the Black Sea.
The authors then investigate patterns of extreme genomic differentiation among populations, and uncover a remarkable pattern of parallel differentiation in these populations. In an outlier scan, Riquet and colleagues find numerous SNPs in one genomic region that separates northern and southern Atlantic populations. Quiet surprisingly, this same genomic region appears to differentiate populations living in marine and lagoon habitats in the Mediterranean. The idea that parallel patterns of genomic differentiation may underlie adaptation to differing environmental scenarios has not yet received much attention. This paper should change that. This paper is particularity impressive in that the authors uncovered this intriguing pattern with under three hundred SNPs. Future genome scale studies will uncover the genomic basis behind this unusual case of parallelism.
References
[1] Riquet, F., Liautard-Haag, C., Woodall, L., Bouza, C., Louisy, P., Hamer, B., Otero-Ferrer, F., Aublanc, P., Béduneau, V., Briard, O., El Ayari, T., Hochscheid, S. Belkhir, K., Arnaud-Haond, S., Gagnaire, P.-A., Bierne, N. (2018). Parallel pattern of differentiation at a genomic island shared between clinal and mosaic hybrid zones in a complex of cryptic seahorse lineages. bioRxiv, 161786, ver. 4 recommended and peer-reviewed by PCI Evol Biol. doi: 10.1101/161786
Reviews: 2
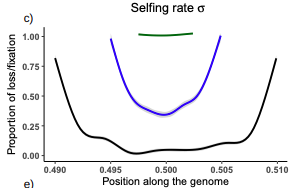
Conditions for maintaining and eroding pseudo-overdominance and its contribution to inbreeding depression
Pseudo-overdominance: how linkage and selection can interact and oppose to purging of deleterious mutations.
Recommended by Sylvain Glémin based on reviews by Yaniv Brandvain, Lei Zhao and 1 anonymous reviewerMost mutations affecting fitness are deleterious and they have many evolutionary consequences. The dynamics and consequences of deleterious mutations are a long-standing question in evolutionary biology and a strong theoretical background has already been developed, for example, to predict the mutation load, inbreeding depression or background selection. One of the classical results is that inbreeding helps purge partially recessive deleterious mutations by exposing them to selection in homozygotes. However, this mainly results from single-locus considerations. When interactions among several, more or less linked, deleterious mutations are taken into account, peculiar dynamics can emerge. One of them, called pseudo-overdominance (POD), corresponds to the maintenance in a population of two (or more) haplotype blocks composed of several recessive deleterious mutations in repulsion that mimics overdominance. Indeed, homozygote individuals for one of the haplotype blocks expose many deleterious mutations to selection whereas they are reciprocally masked in heterozygotes, leading to higher fitness of heterozygotes compared to both homozygotes. A related process, called associative overdominance (AOD) is the effect of such deleterious alleles in repulsion on the linked neutral variation that can be increased by AOD. Although this possibility has been recognized for a long time (Otha and Kimura 1969), it has been mainly considered an anecdotal process. Recently, both theoretical (Zhao and Charlesworth 2016) and genomic analyses (Gilbert et al. 2020) have renewed interest in such a process, suggesting that it could be important in weakly recombining regions of a genome. Donald Waller (2021) - one of the co-authors of the current work - also recently proposed that POD could be quantitatively important with broad implications, and could resolve some unexplained observations such as the maintenance of inbreeding depression in highly selfing species. Yet, a proper theoretical framework analysing the effect of inbreeding on POD was lacking.
In this theoretical work, Diala Abu Awad and Donald Waller (2022) addressed this question through an elegant combination of analytical predictions and intensive multilocus simulations. They determined the conditions under which POD can be maintained and how long it could resist erosion by recombination, which removes the negative association between deleterious alleles (repulsion) at the core of the mechanism. They showed that under tight linkage, POD regions can persist for a long time and generate substantial segregating load and inbreeding depression, even under inbreeding, so opposing (for a while) to the purging effect. They also showed that background selection can affect the genomic structure of POD regions by rapidly erasing weak POD regions but maintaining strong POD regions (i.e with many tightly linked deleterious alleles).
These results have several implications. They can explain the maintenance of inbreeding depression despite inbreeding (as anticipated by Waller 2021), which has implications for the evolution of mating systems. If POD can hardly emerge under high selfing, it can persist from an outcrossing ancestor long after the transition towards a higher selfing rate and could explain the maintenance of mixed mating systems(which is possible with true overdominance, see Uyenoyama and Waller 1991). The results also have implications for genomic analyses, pointing to regions of low or no recombination where POD could be maintained, generating both higher diversity and heterozygosity than expected and variance in fitness. As structural variations are likely widespread in genomes with possible effects on suppressing recombination (Mérot et al. 2020), POD regions should be checked more carefully in genomic analyses (see also Gilbert et al. 2020).
Overall, this work should stimulate new theoretical and empirical studies, especially to assess how quantitatively strong and widespread POD can be. It also stresses the importance of properly considering genetic linkage genome-wide, and so the role of recombination landscapes in determining patterns of diversity and fitness effects.
Awad DA, Waller D (2022) Conditions for maintaining and eroding pseudo-overdominance and its contribution to inbreeding depression. bioRxiv, 2021.12.16.473022, ver. 3 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2021.12.16.473022
Gilbert KJ, Pouyet F, Excoffier L, Peischl S (2020) Transition from Background Selection to Associative Overdominance Promotes Diversity in Regions of Low Recombination. Current Biology, 30, 101-107.e3. https://doi.org/10.1016/j.cub.2019.11.063
Mérot C, Oomen RA, Tigano A, Wellenreuther M (2020) A Roadmap for Understanding the Evolutionary Significance of Structural Genomic Variation. Trends in Ecology & Evolution, 35, 561–572. https://doi.org/10.1016/j.tree.2020.03.002
Ohta T, Kimura M (1969) Linkage disequilibrium at steady state determined by random genetic drift and recurrent mutation. Genetics, 63, 229–238. https://doi.org/10.1093/genetics/63.1.229
Uyenoyama MK, Waller DM (1991) Coevolution of self-fertilization and inbreeding depression II. Symmetric overdominance in viability. Theoretical Population Biology, 40, 47–77. https://doi.org/10.1016/0040-5809(91)90046-I
Waller DM (2021) Addressing Darwin’s dilemma: Can pseudo-overdominance explain persistent inbreeding depression and load? Evolution, 75, 779–793. https://doi.org/10.1111/evo.14189
Zhao L, Charlesworth B (2016) Resolving the Conflict Between Associative Overdominance and Background Selection. Genetics, 203, 1315–1334. https://doi.org/10.1534/genetics.116.188912

Variation in competitive ability with mating system, ploidy and range expansion in four Capsella species
When ecology meets genetics: Towards an integrated understanding of mating system transitions and diversity
Recommended by Sylvain Billiard and Henrique Teotonio based on reviews by Yaniv Brandvain, Henrique Teotonio and 1 anonymous reviewerIn the 19th century, C. Darwin and F. Delpino engaged in a debate about the success of species with different reproduction modes, with the later favouring the idea that monoecious plants capable of autonomous selfing could spread more easily than dioecious plants (or self-incompatible hermaphroditic plants) if cross-pollination opportunities were limited [1]. Since then, debate has never faded about how natural selection is responsible for transitions to selfing and can explain the diversity and distribution of reproduction modes we observe in the natural world [2, 3].
Explanations for mating systems diversity, and transitions to selfing in particular, generally fall into two categories: either genetic or ecological. On the genetic side, many theoretical works showed a critical role for mutation load and inbreeding depression, transmission advantage and reproductive assurance in the evolution of selfing, e.g. [4]. Many experimental works were conducted to test theoretical hypotheses and predictions, especially regarding the magnitude of inbreeding depression; see [5] for a review. Ecologically, the presence of selfing populations is usually correlated with fragmented and harsh habitats, on the periphery of ancestral outcrossing populations. The cause of this distribution could be that selfers are better dispersers and colonizers than outcrossers, or variations in other life-history traits [6]. Yet, few experiments were run to assess whether selfing species or populations have effectively different ecological characteristics, and even scarcer are experiments evaluating both the roles of mutational load and life-history traits evolution. This is the aim of the present study by X. Yang et al [7].
The study of Yang et al [7], together with that of Petrone Mendoza et al. [8], supervised by S. Glémin and M. Lascoux, is probably one of the first to conduct experiments where the competitive abilities are compared between and within species. Using 4 species of the Capsella genus, annual plants from the mustard family, they tested the theoretical predictions that i) the transition from outcrossing to selfing resulted in reduced competitive ability at higher densities, because of the accumulation of deleterious mutations and/or the evolution of life-history traits in an open habitat and a colonization/dispersal trade-off; ii) that reduced competitive ability of selfers should be less pronounced in polyploid then diploid species because the effect of partially recessive deleterious mutations would be buffered; and iii) that competitive ability of selfers should decline with historical range expansion because of the expansion load [9].
Of the 4 Capsella species studied, only one of them, presumably the ancestral, is a diploid outcrosser with a small distribution but large population sizes. The three other species are selfers, two diploids with independent histories of transitions from outcrossing, and another, tetraploid, resulting from a recent hybridization between one of the diploid selfer and the diploid outcrossing ancestor. Many accessions from each species were sampled and individuals assayed for their competitive ability against a tester species or alone, for vegetative and reproductive traits. The measured vegetative traits (rosette surface at two stages, growth rate and flowering probability) showed no differentiation between selfers and outcrossers. To the contrary, reproductive traits (number of flowers) followed theoretical predictions: selfing species are more sensitive to competition than the outcrossing species, with polyploid selfing species being intermediate between the diploid selfers and the diploid outcrosser, and within the tetraploid selfing species (where sampling was quite significant across a large geographical range) sensitivity to competition increased with range expansion.
The study of Yang et al. [7] suffers from several limitations, such that alternative explanations cannot be discarded in the absence of further experimental data. They nonetheless provide the reader with a nice discussion and prospects on how to untwine the causes and the consequences of transitions to selfing. Their study also brings up to date questions about the joint evolution of mating system and life-history traits, which needs a renewed interest from an empirical and theoretical point of view. The results of Yang et al. raise for instance the question of whether it is indeed expected that only reproductive traits, and not vegetative traits, should evolve with the transition to selfing.
The recommandation and evaluation of this paper have been made in collaboration with Thomas Lesaffre.
References
[1] Darwin, C. R. (1876). The effects of cross and self fertilization in the vegetable kingdom. London: Murray.
[2] Stebbins, G. L. (1957). Self fertilization and population variability in the higher plants. The American Naturalist, 91, 337-354. doi: 10.1086/281999
[3] Harder, L.D. & Barrett, S. C. H. (2006). Ecology and evolution of flowers. Oxford: Oxford University Press.
[4] Porcher, E. & Lande, R. (2005). The evolution of self-fertilization and inbreeding depression under pollen discounting and pollen limitation. Journal of Evolutionary Biology, 18(3), 497-508. doi: 10.1111/j.1420-9101.2005.00905.x
[5] Winn, A.A., et al. (2011). Analysis of inbreeding depression in mixed-mating plants provides evidence for selective interference and stable mixed mating. Evolution, 65(12), 3339-3359. doi: 10.1111/j.1558-5646.2011.01462.x
[6] Munoz, F., Violle, C. & Cheptou, P.-O. (2016). CSR ecological strategies and plant mating systems: outcrossing increases with competitiveness but stress-tolerance is related to mixed mating. Oikos, 125(9), 1296-1303. doi: 10.1111/oik.02328
[7] Yang, X., Lascoux, M. & Glémin, S (2018). Variation in competitive ability with mating system, ploidy and range expansion in four Capsella species. bioRxiv, 214866, ver. 5 recommended and peer-reviewed by PCI Evol Biol. doi: 10.1101/214866
[8] Petrone Mendoza, S., Lascoux, M. & Glémin, S. (2018). Competitive ability of Capsella species with different mating systems and ploidy levels. Annals of Botany 121(6), 1257-1264. doi: 10.1093/aob/mcy014
[9] Peischl, S. & Excoffier, L. (2015). Expansion load: recessive mutations and the role of standing genetic variation. Molecular Ecology, 24(9): 2084-2094. doi: 10.1111/mec.13154