
LASCOUX Martin
- Ecology and Genetics, Uppsala University, Uppsala, Sweden
- Adaptation, Evolutionary Applications, Genotype-Phenotype, Molecular Evolution, Phylogeography & Biogeography, Population Genetics / Genomics
- recommender
Recommendation: 1
Reviews: 0
Recommendation: 1
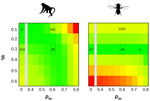
How much does Ne vary among species?
Further questions on the meaning of effective population size
Recommended by Martin Lascoux based on reviews by 3 anonymous reviewersIn spite of its name, the effective population size, Ne, has a complex and often distant relationship to census population size, as we usually understand it. In truth, it is primarily an abstract concept aimed at measuring the amount of genetic drift occurring in a population at any given time. The standard way to model random genetic drift in population genetics is the Wright-Fisher model and, with a few exceptions, definitions of the effective population size stems from it: “a certain model has effective population size, Ne, if some characteristic of the model has the same value as the corresponding characteristic for the simple Wright-Fisher model whose actual size is Ne” (Ewens 2004). Since Sewall Wright introduced the concept of effective population size in 1931 (Wright 1931), it has flourished and there are today numerous definitions of it depending on the process being examined (genetic diversity, loss of alleles, efficacy of selection) and the characteristic of the model that is considered. These different definitions of the effective population size were generally introduced to address specific aspects of the evolutionary process. One aspect that has been hotly debated since the first estimates of genetic diversity in natural populations were published is the so-called Lewontin’s paradox (1974). Lewontin noted that the observed variation in heterozygosity across species was much smaller than one would expect from the neutral expectations calculated with the actual size of the species.
In essence, what Galtier and Rousselle propose in their clever paper is to introduce a new approach to compare effective population sizes across species and thereby a new way to address Lewontin’s paradox. Classically, the effective population size in this type of comparative genomic studies is simply estimated from nucleotide diversity at putatively neutral sites using the equation relating levels of diversity (θ) to mutation rate per generation (μ) and effective population size, Ne, θ = 4Neμ. As Galtier and Rousselle point out there are many issues with this approach. In particular, although we can now estimate θ very precisely, we generally do not have a reliable estimate of the mutation rate, and the method rests on many, unwarranted, assumptions; for example that the population is at mutation-drift equilibrium. Instead they propose to estimate the effective population size from the load of segregating deleterious mutations which can be summarized by the ratio of nonsynonymous to synonymous mutations, πN/πS: small-Ne species are expected to accumulate more deleterious mutations and carry a higher load than large-Ne ones at selection/drift equilibrium (Ohta et al. 1973; Welch et al. 2008). At first glance, this suggestion seems counterintuitive since considering sites under selection undoubtedly adds a new layer of complexity to an already intricate situation. Indeed, one is now bringing to the brew another elusive object, namely, the Distribution of Fitness Effect of mutations (DFE). However, estimating Ne from the load of segregating deleterious mutations may actually simplify the situation in two important ways. First, using πN/πS does not require assumption about μ (as the mutation rate will cancel out in the ratio). Second, the ratio of nonsynonymous to synonymous nucleotide diversity, reaches equilibrium faster than the nucleotide diversity at synonymous site after a change in population size, so we can hope for less sensitivity to (often unknown) recent demographic history (Brandvain and Wright 2016).
Extending recent developments in the estimation of the DFE, Galtier and Rousselle eventually obtain estimates of the average deleterious effect, 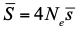 , where Ne is the effective population size and
, where Ne is the effective population size and  is the mean fitness effect of non-synonymous mutations. Assuming further that distinct species share a common DFE and therefore a common
is the mean fitness effect of non-synonymous mutations. Assuming further that distinct species share a common DFE and therefore a common  , they obtain estimates of the between species ratio of Ne from the between species ratio of
, they obtain estimates of the between species ratio of Ne from the between species ratio of  . Applying their newly developed approach to various datasets they conclude that the power of drift varies by a factor of at least 500 between large-Ne (Drosophila) and small-Ne species (H. sapiens). This is an order of magnitude larger than what would be obtained by comparing estimates of the variation in neutral diversity. Hence the proposed approach seems to have gone some way in making Lewontin’s paradox less paradoxical. But, perhaps more importantly, as the authors tersely point out at the end of the abstract their results further questions the meaning of Ne parameters in population genetics. And arguably this could well be the most important contribution of their study and something that is badly needed.
. Applying their newly developed approach to various datasets they conclude that the power of drift varies by a factor of at least 500 between large-Ne (Drosophila) and small-Ne species (H. sapiens). This is an order of magnitude larger than what would be obtained by comparing estimates of the variation in neutral diversity. Hence the proposed approach seems to have gone some way in making Lewontin’s paradox less paradoxical. But, perhaps more importantly, as the authors tersely point out at the end of the abstract their results further questions the meaning of Ne parameters in population genetics. And arguably this could well be the most important contribution of their study and something that is badly needed.
References
Brandvain Y, Wright SI (2016) The Limits of Natural Selection in a Nonequilibrium World. Trends in Genetics, 32, 201–210. doi: 10.1016/j.tig.2016.01.004
Ewens WJ (2010) Mathematical Population Genetics: Theoretical Introduction. Springer-Verlag New York Inc., New York, NY. doi: 10.1007/978-0-387-21822-9
Galtier N, Rousselle M (2020) How much does Ne vary among species? bioRxiv, 861849, ver. 4 peer-reviewed and recommended by PCI Evolutionary Biology. doi: 10.1101/861849
Lewontin RC (1974) The genetic basis of evolutionary change. Columbia University Press, New York.
Ohta T (1973) Slightly Deleterious Mutant Substitutions in Evolution. Nature, 246, 96–98. doi: 10.1038/246096a0
Welch JJ, Eyre-Walker A, Waxman D (2008) Divergence and Polymorphism Under the Nearly Neutral Theory of Molecular Evolution. Journal of Molecular Evolution, 67, 418–426. doi: 10.1007/s00239-008-9146-9
Wright S (1931) Evolution in Mendelian Populations. Genetics, 16, 97–159. url: https://www.genetics.org/content/16/2/97