
FRAGATA Inês
- MITE2: Multidisciplinary Investigation Targeting Ecology and Evolution, cE3c/FCUL, Lisbon, Portugal
- Adaptation, Evolutionary Dynamics, Evolutionary Ecology, Experimental Evolution, Genotype-Phenotype, Life History, Molecular Evolution, Phenotypic Plasticity, Population Genetics / Genomics, Species interactions
- recommender, manager
Recommendations: 5
Reviews: 0
Recommendations: 5

Evolutionary responses of energy metabolism, development, and reproduction to artificial selection for increasing heat tolerance in Drosophila subobscura
The other side of the evolution of heat tolerance: correlated responses in metabolism and life-history traits
Recommended by Inês Fragata and Pedro Simões based on reviews by Marija Savić Veselinović and 1 anonymous reviewerUnderstanding how species respond to environmental changes is becoming increasingly important in order to predict the future of biodiversity and species distributions under current global warming conditions (Rezende 2020; Bennett et al 2021). Two key factors to take into account in these predictions are the tolerance of organisms to heat stress and subsequently how they adapt to increasingly warmer temperatures. Coupled with this, one important factor that is often overlooked when addressing the evolution of thermal tolerance, is the correlated responses in traits that are important to fitness, such as life histories, behavior and the underlying metabolic processes.
The rate and intensity of the thermal stress are expected to be major factors in shaping the evolution of heat tolerance and correlated responses in other traits. For instance, lower rates of thermal stress are predicted to select for individuals with a slower metabolism (Santos et al 2012), whereas low metabolism is expected to lead to a lower reproductive rate (Dammhahn et al 2018). To quantify the importance of the rate and intensity of thermal stress on the evolutionary response of heat tolerance and correlated response in behavior, Mesas et al (2021) performed experimental evolution in Drosophila subobscura using selective regimes with slow or fast ramping protocols. Whereas both regimes showed increased heat tolerance with similar evolutionary rates, the correlated responses in thermal performance curves for locomotor behavior differed between selection regimes. These findings suggest that thermal rate and intensity may shape the evolution of correlated responses in other traits, urging the need to understand possible correlated responses at relevant levels such as life history and metabolism.
In the present contribution, Mesas and Castañeda (2022) investigate whether the disparity in thermal performance curves observed in the previous experiment (Mesas et al 2021) could be explained by differences in metabolic energy production and consumption, and how this correlated with the reproductive output (fecundity and viability). Overall, the authors show some evidence for lowered enzyme activity and increased performance in life-history traits, particularly for the slow-ramping selected flies. Specifically, the authors observe a reduction in glucose metabolism and increased viability when evolving under slow ramping stress. Interestingly, both regimes show a general increase in fecundity, suggesting that adaptation to these higher temperatures is not costly (for reproduction) in the ancestral environment. The evidence for a somewhat lower metabolism in the slow-ramping lines suggests the evolution of a slow “pace of life”. The “pace of life” concept tries to bridge variation across several levels namely metabolism, physiology, behavior and life history, with low “pace of life” organisms presenting lower metabolic rates, later reproduction and higher longevity than fast “pace of life” organisms (Dammhahn et al 2018, Tuzun & Stocks 2022). As the authors state there is not a clear-cut association with the expectations of the pace of life hypothesis since there was evidence for increased reproductive output under both selection intensity regimes. This suggests that, given sufficient trait genetic variance, positively correlated responses may emerge during some stages of thermal evolution. As fecundity estimates in this study were focussed on early life, the possibility of a decrease in the cumulative reproductive output of the selected flies, even under benign conditions, cannot be excluded. This would help explain the apparent paradox of increased fecundity in selected lines. In this context, it would also be interesting to explore the variation in reproductive output at different temperatures, i.e to obtain thermal performance curves for life histories.
Mesas and Castañeda (2022) raise important questions to pursue in the future and contribute to the growing evidence that, in order to predict the distribution of ectothermic species under current global warming conditions, we need to expand beyond determining the physiological thermal limits of each organism (Parratt et al 2021). Ultimately, integrating metabolic, life-history and behavioral changes during evolution under different thermal stresses within a coherent framework is key to developing better predictions of temperature effects on natural populations.
References
Bennett, J.M., Sunday, J., Calosi, P. et al. The evolution of critical thermal limits of life on Earth. Nat Commun 12, 1198 (2021). https://doi.org/10.1038/s41467-021-21263-8
Dammhahn, M., Dingemanse, N.J., Niemelä, P.T. et al. Pace-of-life syndromes: a framework for the adaptive integration of behaviour, physiology and life history. Behav Ecol Sociobiol 72, 62 (2018). https://doi.org/10.1007/s00265-018-2473-y
Mesas, A, Jaramillo, A, Castañeda, LE. Experimental evolution on heat tolerance and thermal performance curves under contrasting thermal selection in Drosophila subobscura. J Evol Biol 34, 767– 778 (2021). https://doi.org/10.1111/jeb.13777
Mesas, A, Castañeda, LE Evolutionary responses of energy metabolism, development, and reproduction to artificial selection for increasing heat tolerance in Drosophila subobscura. bioRxiv, 2022.02.03.479001, ver. 4 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2022.02.03.479001
Parratt, S.R., Walsh, B.S., Metelmann, S. et al. Temperatures that sterilize males better match global species distributions than lethal temperatures. Nat. Clim. Chang. 11, 481–484 (2021). https://doi.org/10.1038/s41558-021-01047-0
Santos, M, Castañeda, LE, Rezende, EL Keeping pace with climate change: what is wrong with the evolutionary potential of upper thermal limits? Ecology and evolution, 2(11), 2866-2880 (2012). https://doi.org/10.1002/ece3.385
Tüzün, N, Stoks, R. A fast pace-of-life is traded off against a high thermal performance. Proceedings of the Royal Society B, 289(1972), 20212414 (2022). https://doi.org/10.1098/rspb.2021.2414
Rezende, EL, Bozinovic, F, Szilágyi, A, Santos, M. Predicting temperature mortality and selection in natural Drosophila populations. Science, 369(6508), 1242-1245 (2020). https://doi.org/10.1126/science.aba9287

Landscape connectivity alters the evolution of density-dependent dispersal during pushed range expansions
Phenotypic evolution during range expansions is contingent on connectivity and density dependence
Recommended by Inês Fragata based on reviews by 3 anonymous reviewersUnderstanding the mechanisms underlying range expansions is key for predicting species distributions in response to environmental changes (such as global warming) and managing the global expansion of invasive species (Parmesan 2006; Suarez & Tsutsui 2008). Traditionally, two types of ecological processes were studied as essential in shaping range expansion: dispersal and population growth. However, ecology and evolution are intertwined in range expansions, as phenotypic evolution of traits involved in demographic and dispersal patterns and processes can affect and be affected by ecological dynamics, representing a full eco-evolutionary loop (Williams et al. 2019; Miller et al. 2020).
Range expansions can be characterized by the type of population growth and dispersal, divided into pushed or pulled range expansions. Species that have high dispersal and high population growth at low densities present pulled range expansions (pulled by individuals from the edge populations). In contrast, populations presenting increased growth rate at intermediate densities (due to Allee effects - Allee & Bowen 1932; i.e. where growth rate decreases at lower densities) and high dispersal at high densities present pushed range expansions (driven by individuals from core and intermediate populations) (Gandhi et al. 2016). Importantly, the type of expansion is expected to have very different consequences on the genetic (and therefore) phenotypic composition of core and edge populations. Specifically, genetic variability is expected to be lower in populations experiencing pulled expansions and higher in populations involved in pushed expansions (Gandhi et al. 2016; Miller et al. 2020). However, it is not always possible to distinguish between pulled and pushed expansions, as variation in speed and shape can overlap between the two types. In addition, it is difficult to experimentally manipulate the strength of the Allee effect to create pushed versus pulled expansions. Thus, several critical predictions regarding the genetic and phenotypic composition of pulled and pushed expansions are lacking empirical tests (but see Gandhi et al. 2016).
In a previous study, Dahirel et al. (2021a) combined simulations and experimental evolution of the small wasps Trichogramma brassicae to show that low connectivity led to more pushed expansions, and higher connectivity generated more pulled expansions. In accordance with theoretical predictions, this led to reduced genetic diversity in pulled expansions, and the reverse pattern in pushed expansions. However, the question of how pulled and pushed expansions affect trait evolution remained unanswered.
In this follow-up study, Dahirel et al. (2021b) tackled this issue and linked the changes in connectivity and type of expansion with the phenotypic evolution of several traits using individuals from their previous experiment. Namely, the authors compared core and edge populations with founder strains to test how evolution in pushed vs. pulled expansions affected wasp size, short movement, fecundity, dispersal, and density dependent dispersal. When density dependence was not accounted for, phenotypic changes in edge populations did not match the expectations from changes in expansion dynamics. This could be due to genetic trade-offs between traits that limit phenotypic evolution (Urquhart & Williams 2021).
However, when accounting for density dependent dispersal, Dahirel et al. (2021b) observed that more connected landscapes (with pulled expansions) showed positive density dispersal in core populations and negative density dispersal in edge populations, similarly to other studies (e.g. Fronhofer et al. 2017). Interestingly, in pushed (with lower connectivity) landscapes, such shift was not observed. Instead, edge populations maintained positive density dispersal even after 14 generations of expansion, whereas core populations showed higher dispersal at lower density. The authors suggest that this seemingly contradictory result is due to a combination of three processes: 1) the expansion reduced positive density dispersal in edge populations; 2) reduced connectivity directly increased dispersal costs, increasing high density dispersal; and 3) reduced connectivity indirectly caused demographic stochasticity (and reduced temporal variability in patches) leading to higher dispersal at low density in core populations. However, these results must be taken with a grain of salt, since only one of the four experimental replicates were used in the density dependent dispersal experiment. In range expansions experiments, replication is fundamental, since stochastic processes (such as gene surfing, where alleles maybe rise in frequency due by chance) are prevalent (Miller et al. 2020), and results are highly dependent on sample size, or number of replicate populations analysed.
Having said that, results from Dahirel et al. (2021b) highlight the importance to contextualize the management of invasions and species distribution, since it is thought that pulled expansions are more prevalent in nature, but pushed expansions can be more important in scenarios where patchiness is high, such as urban landscapes. Moreover, Dahirel's et al. (2021b) study is a first step showing that accounting for trait density dependence is crucial when following phenotypic evolution during range expansion, and that evolution of density dependent traits may be constrained by landscape conditions. This highlights the need to account for both connectivity and density dependence to draw more accurate predictions on the evolutionary and ecological outcomes of range expansions.
References
Allee WC, Bowen ES (1932) Studies in animal aggregations: Mass protection against colloidal silver among goldfishes. Journal of Experimental Zoology, 61, 185–207. https://doi.org/10.1002/jez.1400610202
Dahirel M, Bertin A, Calcagno V, Duraj C, Fellous S, Groussier G, Lombaert E, Mailleret L, Marchand A, Vercken E (2021a) Landscape connectivity alters the evolution of density-dependent dispersal during pushed range expansions. bioRxiv, 2021.03.03.433752, ver. 4 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2021.03.03.433752
Dahirel M, Bertin A, Haond M, Blin A, Lombaert E, Calcagno V, Fellous S, Mailleret L, Malausa T, Vercken E (2021b) Shifts from pulled to pushed range expansions caused by reduction of landscape connectivity. Oikos, 130, 708–724. https://doi.org/10.1111/oik.08278
Fronhofer EA, Gut S, Altermatt F (2017) Evolution of density-dependent movement during experimental range expansions. Journal of Evolutionary Biology, 30, 2165–2176. https://doi.org/10.1111/jeb.13182
Gandhi SR, Yurtsev EA, Korolev KS, Gore J (2016) Range expansions transition from pulled to pushed waves as growth becomes more cooperative in an experimental microbial population. Proceedings of the National Academy of Sciences, 113, 6922–6927. https://doi.org/10.1073/pnas.1521056113
Miller TEX, Angert AL, Brown CD, Lee-Yaw JA, Lewis M, Lutscher F, Marculis NG, Melbourne BA, Shaw AK, Szűcs M, Tabares O, Usui T, Weiss-Lehman C, Williams JL (2020) Eco-evolutionary dynamics of range expansion. Ecology, 101, e03139. https://doi.org/10.1002/ecy.3139
Parmesan C (2006) Ecological and Evolutionary Responses to Recent Climate Change. Annual Review of Ecology, Evolution, and Systematics, 37, 637–669. https://doi.org/10.1146/annurev.ecolsys.37.091305.110100
Suarez AV, Tsutsui ND (2008) The evolutionary consequences of biological invasions. Molecular Ecology, 17, 351–360. https://doi.org/10.1111/j.1365-294X.2007.03456.x
Urquhart CA, Williams JL (2021) Trait correlations and landscape fragmentation jointly alter expansion speed via evolution at the leading edge in simulated range expansions. Theoretical Ecology. https://doi.org/10.1007/s12080-021-00503-z
Williams JL, Hufbauer RA, Miller TEX (2019) How Evolution Modifies the Variability of Range Expansion. Trends in Ecology & Evolution, 34, 903–913. https://doi.org/10.1016/j.tree.2019.05.012

How do invasion syndromes evolve? An experimental evolution approach using the ladybird Harmonia axyridis
Selection on a single trait does not recapitulate the evolution of life-history traits seen during an invasion
Recommended by Inês Fragata and Ben Phillips based on reviews by 2 anonymous reviewersBiological invasions are natural experiments, and often show that evolution can affect dynamics in important ways [1-3]. While we often think of invasions as a conservation problem stemming from anthropogenic introductions [4,5], biological invasions are much more commonplace than this, including phenomena as diverse as natural range shifts, the spread of novel pathogens, and the growth of tumors. A major question across all these settings is which set of traits determine the ability of a population to invade new space [6,7]. Traits such as: increased growth or reproductive rate, dispersal ability and ability to defend from predation often show large evolutionary shifts across invasion history [1,6,8]. Are such multi-trait shifts driven by selection on multiple traits, or a correlated response by multiple traits to selection on one? Resolving this question is important for both theoretical and practical reasons [9,10]. But despite the importance of this issue, it is not easy to perform the necessary manipulative experiments [9].
Foucaud et al. [11] tackled this issue by performing experimental evolution on source populations of the invasive ladybug Harmonia axyridis. The authors tested if selection on a single trait could generate correlated responses in other life history traits. Specifically, they used experimental evolution to impose divergent selection on female mass, and reproductive timing. After ten generations, they found that selection for weight did not affect almost any other life history trait. However, nine generations of selection for faster reproduction led to correlated phenotypic changes in developmental, reproduction and survival rate of populations, although not always in the direction we might have expected. Despite this correlated response, none of their selected lines were able to fully recapitulate the trait shifts seen in natural invasions of this species. This implies that selection during natural invasions is operating on multiple traits; a finding in agreement with our growing understanding of how selection acts during introduction and invasion [12,13].
Populations undergoing a colonization process may also be subject to a multitude of different selective pressures [14,15]. The authors expanded their work in this direction by testing whether food availability alters the observed correlations between life history traits. The pervasiveness of genotype by environment interactions observed also points to a role for multiple selective pressures in shaping the suite of life-history shifts observed in wild ladybug populations. The work from Foucaud and colleagues [11] adds to a small but growing list of important studies that use experimental evolution to investigate how life-history traits evolve, and how they evolve during invasions in particular.
References
[1] Sakai, A.K., Allendorf, F.W., Holt, J.S. et al. (2001). The population biology of invasive species. Annual review of ecology and systematics, 32(1), 305-332. doi: 10.1146/annurev.ecolsys.32.081501.114037
[2] Hairston Jr, N. G., Ellner, S. P., Geber, M. A., Yoshida, T. and Fox, J. A. (2005). Rapid evolution and the convergence of ecological and evolutionary time. Ecology letters, 8(10), 1114-1127. doi: 10.1111/j.1461-0248.2005.00812.x
[3] Chuang, A. and Peterson, C. R. (2016). Expanding population edges: theories, traits, and trade‐offs. Global change biology, 22(2), 494-512. doi: 10.1111/gcb.13107
[4] Whitney, K. D. and Gabler, C. A. (2008). Rapid evolution in introduced species,‘invasive traits’ and recipient communities: challenges for predicting invasive potential. Diversity and Distributions, 14(4), 569-580. doi: 10.1111/j.1472-4642.2008.00473.x
[5] Catullo, R. A., Llewelyn, J., Phillips, B. L. and Moritz, C. C. (2019). The Potential for Rapid Evolution under Anthropogenic Climate Change. Current Biology, 29(19), R996-R1007. doi: 10.1016/j.cub.2019.08.028
[6] Suarez, A. V. and Tsutsui, N. D. (2008). The evolutionary consequences of biological invasions. Molecular Ecology, 17(1), 351-360. doi: 10.1111/j.1365-294X.2007.03456.x
[7] Deforet, M., Carmona-Fontaine, C., Korolev, K. S. and Xavier, J. B. (2019). Evolution at the edge of expanding populations. The American Naturalist, 194(3), 291-305. doi: 10.1086/704594
[8] Phillips, B. L., Brown, G. P., and Shine, R. (2010). Life‐history evolution in range‐shifting populations. Ecology, 91(6), 1617-1627. doi: 10.1890/09-0910.1
[9] Colautti, R. I. and Lau, J. A. (2015). Contemporary evolution during invasion: evidence for differentiation, natural selection, and local adaptation. Molecular ecology, 24(9), 1999-2017. doi: 10.1111/mec.13162
[10] Szűcs, M., Melbourne, B. A., Tuff, T., Weiss‐Lehman, C. and Hufbauer, R. A. (2017). Genetic and demographic founder effects have long‐term fitness consequences for colonising populations. Ecology Letters, 20(4), 436-444. doi: 10.1111/ele.12743
[11] Foucaud, J., Hufbauer, R. A., Ravigné, V., Olazcuaga, L., Loiseau, A., Ausset, A., Wang, S., Zang, L.-S., Lemenager, N., Tayeh, A., Weyna, A., Gneux, P., Bonnet, E., Dreuilhe, V., Poutout, B., Estoup, A. and Facon, B. (2020). How do invasion syndromes evolve? An experimental evolution approach using the ladybird Harmonia axyridis. bioRxiv, 849968 ver. 4 peer-reviewed and recommended by PCI Evolutionary Biology. doi: 10.1101/849968
[12] Simons, A. M. (2003). Invasive aliens and sampling bias. Ecology Letters, 6(4), 278-280. doi: 10.1046/j.1461-0248.2003.00430.x
[13] Phillips, B. L. and Perkins, T. A. (2019). Spatial sorting as the spatial analogue of natural selection. Theoretical Ecology, 12(2), 155-163. doi: 10.1007/s12080-019-0412-9
[14] Lavergne, S. and Molofsky, J. (2007). Increased genetic variation and evolutionary potential drive the success of an invasive grass. Proceedings of the National Academy of Sciences, 104(10), 3883-3888. doi: 10.1073/pnas.0607324104
[15] Moran, E. V. and Alexander, J. M. (2014). Evolutionary responses to global change: lessons from invasive species. Ecology Letters, 17(5), 637-649. doi: 10.1111/ele.12262
Cost of resistance: an unreasonably expensive concept
Let’s move beyond costs of resistance!
Recommended by Inês Fragata and Claudia Bank based on reviews by Danna Gifford, Helen Alexander and 1 anonymous reviewerThe increase in the prevalence of (antibiotic) resistance has become a major global health concern and is an excellent example of the impact of real-time evolution on human society. This has led to a boom of studies that investigate the mechanisms and factors involved in the evolution of resistance, and to the spread of the concept of "costs of resistance". This concept refers to the relative fitness disadvantage of a drug-resistant genotype compared to a non-resistant reference genotype in the ancestral (untreated) environment.
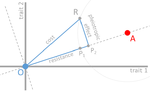
In their paper, Lenormand et al. [1] discuss the history of this concept and highlight its caveats and limitations. The authors address both practical and theoretical problems that arise from the simplistic view of "costly resistance" and argue that they can be prejudicial for antibiotic resistance studies. For a better understanding, they visualize their points of critique by means of Fisher's Geometric model.
The authors give an interesting historical overview of how the concept arose and speculate that it emerged (during the 1980s) in an attempt by ecologists to spread awareness that fitness can be environment-dependent, and because of the concept's parallels to trade-offs in life-history evolution. They then identify several problems that arise from the concept, which, besides the conceptual misunderstandings that they can cause, are important to keep in mind when designing experimental studies.
The authors highlight and explain the following points:
1. Costs of resistance do not necessarily imply pleiotropic effects of a resistance mutation, and pleiotropy is not necessarily the cause of fitness trade-offs.
2. Any non-treated environment and any treatment dose can result in a different cost.
3. Different reference genotypes may result in different costs. Specifically, the reference genotype has to be "optimally" adapted to the reference environment to provide an accurate measurement of costs.
Lenormand et al.'s paper [1] is a timely perspective piece in light of the ever-increasing efforts to understand and tackle resistance evolution [2]. Although some readers may shy away from the rather theoretical presentation of the different points of concern, it will be useful for both theoretical and empirical readers by illustrating the misconceptions that can arise from the concept of the cost of resistance. Ultimately, the main lesson to be learned from this paper may not be to ban the term "cost of resistance" from one's vocabulary, but rather to realize that the successful fight against drug resistance requires more differential information than the measurement of fitness effects in a drug-treated vs. non-treated environment in the lab [3-4]. Specifically, a better integration of the ecological aspects of drug resistance evolution and maintenance is needed [5], and we are far from a general understanding of how environmental factors interact and influence an organism's (absolute and relative) fitness and the effect of resistance mutations.
References
[1] Lenormand T, Harmand N, Gallet R. 2018. Cost of resistance: an unreasonably expensive concept. bioRxiv 276675, ver. 3 peer-reviewed by Peer Community In Evolutionary Biology. doi: 10.1101/276675
[2] Andersson DI and Hughes D. Persistence of antibiotic resistance in bacterial populations. 2011. FEMS Microbiology Reviews, 35: 901-911. doi: 10.1111/j.1574-6976.2011.00289.x
[3] Chevereau G, Dravecká M, Batur T, Guvenek A, Ayhan DH, Toprak E, Bollenbach T. 2015. Quantifying the determinants of evolutionary dynamics leading to drug resistance. PLoS biology 13, e1002299. doi: 10.1371/journal.pbio.1002299
[4] Bengtsson-Palme J, Kristiansson E, Larsson DGJ. 2018. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiology Reviews 42: 68–80. doi: 10.1093/femsre/fux053
[5] Hiltunen T, Virta M, Laine AL. 2017. Antibiotic resistance in the wild: an eco-evolutionary perspective. Philosophical Transactions of the Royal Society B: Biological Sciences 372: 20160039. doi: 10.1098/rstb.2016.0039
Fisher's geometrical model and the mutational patterns of antibiotic resistance across dose gradients
What doesn’t kill us makes us stronger: can Fisher’s Geometric model predict antibiotic resistance evolution?
Recommended by Inês Fragata and Claudia BankThe increasing number of reported cases of antibiotic resistance is one of today’s major public health concerns. Dealing with this threat involves understanding what drives the evolution of antibiotic resistance and investigating whether we can predict (and subsequently avoid or circumvent) it [1].
One of the most illustrative and common models of adaptation (and, hence, resistance evolution) is Fisher’s Geometric Model (FGM). The original model maps phenotypes to fitness, meaning that each point in the fitness landscape corresponds to a phenotype rather than a genotype. However, it has been shown that when mutations are numerous enough, FGM can also describe adaptive walks in genotype space [2]. Nevertheless, limitations have been highlighted, particularly when trying to study complex scenarios such as antibiotic resistance evolution [3].
Harmand et al. [4] incorporated three extensions to the FGM, which allowed them to match the mutational patterns of antibiotic resistance that they obtained from a screen across a gradient of drug concentrations. The implemented extensions took into account that: 1) only a subset of mutations may contribute to traits under selection, reflecting that not all regions in the genome affect the ability to resist antibiotics; 2) mutations that confer a fitness increase in one environment may not reflect a similar increase in others, if the selective constraints are different; and 3) different antibiotic concentrations may either constrain the maximum fitness that populations can reach (changing the height of the fitness peak) or change the rate of fitness increase with each mutation (changing the width/slope of the peak).
Traditionally, most empirical fitness landscape studies have focused on a subset of mutations obtained after laboratory evolution in specific conditions [5, 6]. The results obtained in Harmand et al. [4] indicate a potential shortcoming of studying these small fitness landscapes: rather than having a constrained evolutionary path to a resistant phenotype, as previously observed, their results suggest that antibiotic resistance can be the product of mutations in different regions of the genome. Returning to the fitness landscape perspective, this indicates that there are many alternative paths that can lead to the evolution of antibiotic resistance.
This comparison points at a difficult challenge when aiming at developing a predictive framework for evolution: real-time experiments may indicate that evolution is likely to take similar and predictable paths because the strongest and most frequent mutations dictate the outcome, whereas systematic screens of mutants potentially indicate several paths, that may, however, not be relevant in nature. Only a combination of different experimental approaches with motivated theory as presented in Harmand et al. [4] will allow for a better understanding of where in this continuum evolution is taking place in nature, and to which degree we are able to interfere with it in order to slow down adaptation.
References
[1] Palmer AC, and Kishony R. 2013. Understanding, predicting and manipulating the genotypic evolution of antibiotic resistance. Nature Review Genetics 14: 243—248. doi: 10.1038/nrg3351
[2] Tenaillon O. 2014. The utility of Fisher’s geometric model in evolutionary genetics. Annual Review of Ecology, Evolution and Systematics 45: 179—201. doi: 10.1146/annurev-ecolsys-120213-091846
[3] Blanquart F and Bataillon T. 2016. Epistasis and the structure of fitness landscapes: are experimental fitness landscapes compatible with Fisher’s geometric model? Genetics 203: 847—862. doi: 10.1534/genetics.115.182691
[4] Harmand N, Gallet R, Jabbour-Zahab R, Martin G and Lenormand T. 2017. Fisher’s geometrical model and the mutational patterns of antibiotic resistance across dose gradients. Evolution 71: 23—37. doi: 10.1111/evo.13111
[5] de Visser, JAGM, and Krug J. 2014. Empirical fitness landscapes and the predictability of evolution. Nature 15: 480—490. doi: 10.1038/nrg3744
[6] Palmer AC, Toprak E, Baym M, Kim S, Veres A, Bershtein S and Kishony R. 2015. Delayed commitment to evolutionary fate in antibiotic resistance fitness landscapes. Nature Communications 6: 1—8. doi: 10.1038/ncomms8385