
SIMÕES Pedro
- Department of Animal Biology, Centre for Ecology, Evolution and Environmental Changes (cE3c) - Faculdade de Ciências, Universidade de Lisboa, Lisbon, Portugal
- Adaptation, Evolutionary Dynamics, Evolutionary Ecology, Experimental Evolution, Life History, Phenotypic Plasticity, Population Genetics / Genomics
- recommender
Recommendations: 2
Reviews: 3
Recommendations: 2
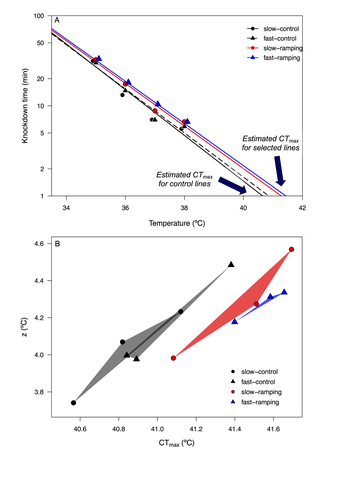
Cross-tolerance evolution is driven by selection of heat tolerance in Drosophila subobscura
Evolution of cross-tolerance: a mechanism to cope with climate change?
Recommended by Pedro Simões based on reviews by Marina Stamenkovic-Radak and 1 anonymous reviewerUnderstanding how populations evolve under thermal stress and how this process shapes the response of other stress responses is an important research topic in the context of thermal adaptation and climate change. In a thermal experimental evolution study in Drosophila subobscura, Castañeda (2024) addressed the correlated responses to selection for increasing knockdown temperature in different resistance traits, either directly related to thermal stress (e.g. knockdown time at different temperatures and CTmax) or not (e.g. desiccation and starvation resistance).
The author found that the evolution of higher knockdown temperature did in fact lead to correlated responses in other stress traits. While such correlations might be expected for the thermal stress traits measured (knockdown time and CTmax), it was perhaps less expectable for desiccation and starvation resistance. However, the general occurrence of correlated evolutionary responses between stressors has been previously described, namely in Drosophila (e.g. see Bubliy and Loeschcke 2005), pointing to a possible genetic link between distinct (thermal) stress traits.
There are however some features that make the findings of this study rather appealing. First, the evidence that the correlated stress responses depend on the intensity of thermal selection (i.e. the warming rate) and on the sex of the organisms. Second, correlated patterns of both desiccation and starvation resistance highlight the possibility of the evolution of a cross-tolerance response, which might positively impact on population ability to evolve under sustained stressful environments (Rodgers and Gomez Izasa 2023). However, it is important to point out that the correlated patterns between these two resistance traits (desiccation and starvation) were not exactly consistent. In fact, the negative correlated response observed for female starvation resistance is thought provoking and argues again a general scenario of cross-tolerance.
While these findings are a step forward for a more multifaceted understanding of thermal adaptation in the context of stressful environments, they also highlight the need for further studies of thermal adaptation namely 1) addressing the underlying physiological and genomic mechanisms that link male and female heat tolerance and the response to other stress resistance traits (namely starvation resistance); 2) testing the extent to which cross-resistance patterns can be generalized to different thermal selection contexts and populations.
In addition, this study also opens new questions considering the scope of correlated evolution to other stress traits, that might be relevant in diverse ecological scenarios. For instance, does selection towards higher heat resistance lead to correlated evolution of cold resistance? And under which circumstances (e.g. different heat selection intensities)? In fact, the occurrence of a positive (or negative) correlation cold and heat stress responses is a topic of high interest, with relevant ecological implications particularly considering the increased thermal fluctuations in natural environments because of climate warming. Cross-tolerance between cold and heat stress responses has been described (Singh 2022, Rodgers and Gomez Izasa 2023). On the other hand, negative correlations (i.e. trade-offs) between these stress traits (Stazione et al. 2020; Schou et al 2022) can impact negatively on populations’ ability to withstand thermal variability.
As climatic changes proceed leading to increasing environmental variability, empirical studies such as that of Castañeda (2024) are critical in the pursue for a multivariate perspective on trait evolution in scenarios of climate change adaptation. Understanding how tolerance to different environmental stressors may evolve and which factors can act as drivers of that variation will ultimately enable better forecasts of climate change effects on biodiversity in nature.
References
Castañeda, LE. Cross-tolerance evolution is driven by selection on heat tolerance in Drosophila subobscura. Biorxiv, ver. 4 peer-reviewed and recommended by Peer Community in Evolutionary Biology (2024). https://www.doi.org/10.1101/2023.09.05.556367
Bubliy, OA, Loeschcke, V. Correlated responses to selection for stress resistance and longevity in a laboratory population of Drosophila melanogaster. J Evol Biol. 18(4):789-803 (2005). https://www.doi.org/10.1111/j.1420-9101.2005.00928.x.
Rodgers, EM, Gomez Isaza, DF. The mechanistic basis and adaptive significance of cross-tolerance: a 'pre-adaptation' to a changing world? J Exp Biol. 226(11):jeb245644 (2023). https://www.doi.org/10.1242/jeb.245644.
Schou, MF, Engelbrecht, A, Brand, Z, Svensson, EI, Cloete, S, Cornwallis, CK. Evolutionary trade-offs between heat and cold tolerance limit responses to fluctuating climates. Sci Adv. 8(21):eabn9580 (2022). https://www.doi.org/10.1126/sciadv.abn9580.
Singh, K, Arun Samant, M, Prasad, NG. Evolution of cross-tolerance in Drosophila melanogaster as a result of increased resistance to cold stress. Sci Rep. 12(1):19536 (2022). https://www.doi.org/10.1038/s41598-022-23674-z.
Stazione, L, Norry, FM, Gomez, FH, Sambucetti, P. Heat knockdown resistance and chill-coma recovery as correlated responses to selection on mating success at high temperature in Drosophila buzzatii. Ecol Evol. 10(4):1998-2006 (2020). https://www.doi.org/10.1002/ece3.6032.

Evolutionary responses of energy metabolism, development, and reproduction to artificial selection for increasing heat tolerance in Drosophila subobscura
The other side of the evolution of heat tolerance: correlated responses in metabolism and life-history traits
Recommended by Inês Fragata and Pedro Simões based on reviews by Marija Savić Veselinović and 1 anonymous reviewerUnderstanding how species respond to environmental changes is becoming increasingly important in order to predict the future of biodiversity and species distributions under current global warming conditions (Rezende 2020; Bennett et al 2021). Two key factors to take into account in these predictions are the tolerance of organisms to heat stress and subsequently how they adapt to increasingly warmer temperatures. Coupled with this, one important factor that is often overlooked when addressing the evolution of thermal tolerance, is the correlated responses in traits that are important to fitness, such as life histories, behavior and the underlying metabolic processes.
The rate and intensity of the thermal stress are expected to be major factors in shaping the evolution of heat tolerance and correlated responses in other traits. For instance, lower rates of thermal stress are predicted to select for individuals with a slower metabolism (Santos et al 2012), whereas low metabolism is expected to lead to a lower reproductive rate (Dammhahn et al 2018). To quantify the importance of the rate and intensity of thermal stress on the evolutionary response of heat tolerance and correlated response in behavior, Mesas et al (2021) performed experimental evolution in Drosophila subobscura using selective regimes with slow or fast ramping protocols. Whereas both regimes showed increased heat tolerance with similar evolutionary rates, the correlated responses in thermal performance curves for locomotor behavior differed between selection regimes. These findings suggest that thermal rate and intensity may shape the evolution of correlated responses in other traits, urging the need to understand possible correlated responses at relevant levels such as life history and metabolism.
In the present contribution, Mesas and Castañeda (2022) investigate whether the disparity in thermal performance curves observed in the previous experiment (Mesas et al 2021) could be explained by differences in metabolic energy production and consumption, and how this correlated with the reproductive output (fecundity and viability). Overall, the authors show some evidence for lowered enzyme activity and increased performance in life-history traits, particularly for the slow-ramping selected flies. Specifically, the authors observe a reduction in glucose metabolism and increased viability when evolving under slow ramping stress. Interestingly, both regimes show a general increase in fecundity, suggesting that adaptation to these higher temperatures is not costly (for reproduction) in the ancestral environment. The evidence for a somewhat lower metabolism in the slow-ramping lines suggests the evolution of a slow “pace of life”. The “pace of life” concept tries to bridge variation across several levels namely metabolism, physiology, behavior and life history, with low “pace of life” organisms presenting lower metabolic rates, later reproduction and higher longevity than fast “pace of life” organisms (Dammhahn et al 2018, Tuzun & Stocks 2022). As the authors state there is not a clear-cut association with the expectations of the pace of life hypothesis since there was evidence for increased reproductive output under both selection intensity regimes. This suggests that, given sufficient trait genetic variance, positively correlated responses may emerge during some stages of thermal evolution. As fecundity estimates in this study were focussed on early life, the possibility of a decrease in the cumulative reproductive output of the selected flies, even under benign conditions, cannot be excluded. This would help explain the apparent paradox of increased fecundity in selected lines. In this context, it would also be interesting to explore the variation in reproductive output at different temperatures, i.e to obtain thermal performance curves for life histories.
Mesas and Castañeda (2022) raise important questions to pursue in the future and contribute to the growing evidence that, in order to predict the distribution of ectothermic species under current global warming conditions, we need to expand beyond determining the physiological thermal limits of each organism (Parratt et al 2021). Ultimately, integrating metabolic, life-history and behavioral changes during evolution under different thermal stresses within a coherent framework is key to developing better predictions of temperature effects on natural populations.
References
Bennett, J.M., Sunday, J., Calosi, P. et al. The evolution of critical thermal limits of life on Earth. Nat Commun 12, 1198 (2021). https://doi.org/10.1038/s41467-021-21263-8
Dammhahn, M., Dingemanse, N.J., Niemelä, P.T. et al. Pace-of-life syndromes: a framework for the adaptive integration of behaviour, physiology and life history. Behav Ecol Sociobiol 72, 62 (2018). https://doi.org/10.1007/s00265-018-2473-y
Mesas, A, Jaramillo, A, Castañeda, LE. Experimental evolution on heat tolerance and thermal performance curves under contrasting thermal selection in Drosophila subobscura. J Evol Biol 34, 767– 778 (2021). https://doi.org/10.1111/jeb.13777
Mesas, A, Castañeda, LE Evolutionary responses of energy metabolism, development, and reproduction to artificial selection for increasing heat tolerance in Drosophila subobscura. bioRxiv, 2022.02.03.479001, ver. 4 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2022.02.03.479001
Parratt, S.R., Walsh, B.S., Metelmann, S. et al. Temperatures that sterilize males better match global species distributions than lethal temperatures. Nat. Clim. Chang. 11, 481–484 (2021). https://doi.org/10.1038/s41558-021-01047-0
Santos, M, Castañeda, LE, Rezende, EL Keeping pace with climate change: what is wrong with the evolutionary potential of upper thermal limits? Ecology and evolution, 2(11), 2866-2880 (2012). https://doi.org/10.1002/ece3.385
Tüzün, N, Stoks, R. A fast pace-of-life is traded off against a high thermal performance. Proceedings of the Royal Society B, 289(1972), 20212414 (2022). https://doi.org/10.1098/rspb.2021.2414
Rezende, EL, Bozinovic, F, Szilágyi, A, Santos, M. Predicting temperature mortality and selection in natural Drosophila populations. Science, 369(6508), 1242-1245 (2020). https://doi.org/10.1126/science.aba9287
Reviews: 3
Phenotypic stasis with genetic divergence
Phenotypic stasis despite genetic divergence and differentiation in Caenorhabditis elegans.
Recommended by Frédéric Guillaume based on reviews by Benoit Pujol and Pedro SimõesExplaining long periods of evolutionary stasis, the absence of change in trait means over geological times, despite the existence of abundant genetic variation in most traits has challenged evolutionary theory since Darwin's theory of evolution by gradual modification (Estes & Arnold 2007). Stasis observed in contemporary populations is even more daunting since ample genetic variation is usually coupled with the detection of selection differentials (Kruuk et al. 2002, Morrissey et al. 2010). Moreover, rapid adaptation to environmental changes in contemporary populations, fuelled by standing genetic variation provides evidence that populations can quickly respond to an adaptive challenge. Explanations for evolutionary stasis usually invoke stabilizing selection as a main actor, whereby optimal trait values remain roughly constant over long periods of time despite small-scale environmental fluctuations. Genetic correlation among traits may also play a significant role in constraining evolutionary changes over long timescales (Schluter 1996). Yet, genetic constraints are rarely so strong as to completely annihilate genetic changes, and they may evolve. Patterns of genetic correlations among traits, as captured in estimates of the G-matrix of additive genetic co-variation, are subject to changes over generations under the action of drift, migration, or selection, among other causes (Arnold et al. 2008). Therefore, under the assumption of stabilizing selection on a set of traits, phenotypic stasis and genetic divergence in patterns of trait correlations may both be observed when selection on trait correlations is weak relative to its effect on trait means.
Mallard et al. (2023) set out to test whether selection or drift may explain the divergence in genetic correlation among traits in experimental lines of the nematode Caenorhabditis elegans and whether stabilizing selection may be a driver of phenotypic stasis. To do so, they analyzed the evolution of locomotion behavior traits over 100 generations of lab evolution in a constant and homogeneous environment after 140 generations of domestication from a largely differentiated set of founder populations. The locomotion traits were transition rates between movement states and direction (still, forward or backward movement). They could estimate the traits' broad-sense G-matrix in three populations at two generations (50 and 100), and in the ancestral mixed population. Similarly, they estimated the shape of the selection surface by regressing locomotion behavior on fertility. Armed with both G-matrix and surface estimates, they could test whether the G's orientation matched selection's orientation and whether changes in G were constrained by selection. They found stasis in trait mean over 100 generations but divergence in the amount and orientation of the genetic variation of the traits relative to the ancestral population. The selected populations changed orientation of their G-matrices and lost genetic variation during the experiment in agreement with a model of genetic drift on quantitative traits. Their estimates of selection also point to mostly stabilizing selection on trait combinations with weak evidence of disruptive selection, suggesting a saddle-shaped selection surface. The evolutionary responses of the experimental populations were mostly consistent with small differentiation in the shape of G-matrices during the 100 generations of stabilizing selection.
Mallard et al. (2023) conclude that phenotypic stasis was maintained by stabilizing selection and drift in their experiment. They argue that their findings are consistent with a "table-top mountain" model of stabilizing selection, whereby the population is allowed some wiggle room around the trait optimum, leaving space for random fluctuations of trait variation, and especially trait co-variation. The model is an interesting solution that might explain how stasis can be maintained over contemporary times while allowing for random differentiation of trait genetic co-variation. Whether such differentiation can then lead to future evolutionary divergence once replicated populations adapt to a new environment is an interesting idea to follow.
References
Arnold, S. J., Bürger, R., Hohenlohe, P. A., Ajie, B. C. and Jones, A. G. 2008. Understanding the evolution and stability of the G-matrix. Evolution 62(10): 2451-2461.
https://doi.org/10.1111/j.1558-5646.2008.00472.x
Estes, S. and Arnold, S. J. 2007. Resolving the Paradox of Stasis: Models with Stabilizing Selection Explain Evolutionary Divergence on All Timescales.. Am. Nat. 169(2): 227-244.
https://doi.org/10.1086/510633
Kruuk, L. E. B., Slate, J., Pemberton, J. M., Brotherstone, S., Guinness, F. and Clutton-Brock, T. 2002. Antler size in red deer: Heritability and selection but no evolution. Evolution 56(8): 1683-1695.
https://doi.org/10.1111/j.0014-3820.2002.tb01480.x
Mallard, F., Noble, L., Guzella, T., Afonso, B., Baer, C. F., Teotónio, H. 2023. Phenotypic stasis with genetic divergence. bioRxiv, ver. 3 peer-reviewed and recommended by Peer Community in Evolutionary Biology.
https://doi.org/10.1101/2022.05.28.493856
Morrissey, M. B., Kruuk, L. E. B. and Wilson, A. J. 2010. The danger of applying the breeder's equation in observational studies of natural populations. J Evolution Biol 23(11): 2277-2288.
https://doi.org/10.1111/j.1420-9101.2010.02084.x
Schluter, D. 1996. Adaptive radiation along genetic lines of least resistance. Evolution 50(5): 1766-1774.
https://doi.org/10.1111/j.1558-5646.1996.tb03563.x
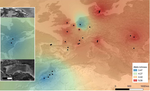

Large-scale geographic survey provides insights into the colonization history of a major aphid pest on its cultivated apple host in Europe, North America and North Africa
The evolutionary puzzle of the host-parasite-endosymbiont Russian doll for apples and aphids
Recommended by Ignacio Bravo based on reviews by Pedro Simões and 1 anonymous reviewerEach individual multicellular organism, each of our bodies, is a small universe. Every living surface -skin, cuticle, bark, mucosa- is the home place to milliards of bacteria, fungi and viruses. They constitute our microbiota. Some of them are essential for certain organisms. Other could not live without their hosts. For many species, the relationship between host and microbiota is so close that their histories are inseparable. The recognition of this biological inextricability has led to the notion of holobiont as the organism ensemble of host and microbiota. When individuals of a particular animal or plant species expand their geographical range, it is the holobiont that expands. And these processes of migration, expansion and colonization are often accompanied by evolutionary and ecological innovations in the interspecies relationships, at the macroscopic level (e.g. novel predator-prey or host-parasite interactions) and at the microscopic level (e.g. changes in the microbiota composition). From the human point of view, these novel interactions can be economically disastrous if they involve and threaten important crop or cattle species. And this is especially worrying in the present context of genetic standardization and intensification for mass-production on the one hand, and of climate change on the other.
With this perspective, the international team led by Amandine Cornille presents a study aiming at understanding the evolutionary history of the rosy apple aphid Dysaphis plantaginea Passerini, a major pest of the cultivated apple tree Malus domestica Borkh (1). The apple tree was probably domesticated in Central Asia, and later disseminated by humans over the world in different waves, and it was probably introduced in Europe by the Greeks. It is however unclear when and where D. plantaginea started parasitizing the cultivated apple tree. The ancestral D. plantaginea could have already infected the wild ancestor of current cultivated apple trees, but the aphid is not common in Central Asia. Alternatively, it may have gained access only later to the plant, possibly via a host jump, from Pyrus to Malus that may have occurred in Asia Minor or in the Caucasus. In the present preprint, Olvera-Vázquez and coworkers have analysed over 650 D. plantaginea colonies from 52 orchards in 13 countries, in Western, Central and Eastern Europe as well as in Morocco and the USA. The authors have analysed the genetic diversity in the sampled aphids, and have characterized as well the composition of the associated endosymbiont bacteria. The analyses detect substantial recent admixture, but allow to identify aphid subpopulations slightly but significantly differentiated and isolated by distance, especially those in Morocco and the USA, as well as to determine the presence of significant gene flow. This process of colonization associated to gene flow is most likely indirectly driven by human interactions. Very interestingly, the data show that this genetic diversity in the aphids is not reflected by a corresponding diversity in the associated microbiota, largely dominated by a few Buchnera aphidicola variants. In order to determine polarity in the evolutionary history of the aphid-tree association, the authors have applied approximate Bayesian computing and machine learning approaches. Albeit promising, the results are not sufficiently robust to assess directionality nor to confidently assess the origin of the crop pest. Despite the large effort here communicated, the authors point to the lack of sufficient data (in terms of aphid isolates), especially originating from Central Asia. Such increased sampling will need to be implemented in the future in order to elucidate not only the origin and the demographic history of the interaction between the cultivated apple tree and the rosy apple aphid. This knowledge is needed to understand how this crop pest struggles with the different seasonal and geographical selection pressures while maintaining high genetic diversity, conspicuous gene flow, differentiated populations and low endosymbiontic diversity.
References
- Olvera-Vazquez SG, Remoué C, Venon A, Rousselet A, Grandcolas O, Azrine M, Momont L, Galan M, Benoit L, David GM, Alhmedi A, Beliën T, Alins G, Franck P, Haddioui A, Jacobsen SK, Andreev R, Simon S, Sigsgaard L, Guibert E, Tournant L, Gazel F, Mody K, Khachtib Y, Roman A, Ursu TM, Zakharov IA, Belcram H, Harry M, Roth M, Simon JC, Oram S, Ricard JM, Agnello A, Beers EH, Engelman J, Balti I, Salhi-Hannachi A, Zhang H, Tu H, Mottet C, Barrès B, Degrave A, Razmjou J, Giraud T, Falque M, Dapena E, Miñarro M, Jardillier L, Deschamps P, Jousselin E, Cornille A (2021) Large-scale geographic survey provides insights into the colonization history of a major aphid pest on its cultivated apple host in Europe, North America and North Africa. bioRxiv, 2020.12.11.421644, ver. 3 peer-reviewed and recommended by Peer Community in Evolutionary Biology. https://doi.org/10.1101/2020.12.11.421644
Environmental specificity in Drosophila-bacteria symbiosis affects host developmental plasticity
Nutrition-dependent effects of gut bacteria on growth plasticity in Drosophila melanogaster
Recommended by Wolf Blanckenhorn based on reviews by Pedro Simões and 1 anonymous reviewerIt is well known that the rearing environment has strong effects on life history and fitness traits of organisms. Microbes are part of every environment and as such likely contribute to such environmental effects. Gut bacteria are a special type of microbe that most animals harbor, and as such they are part of most animals’ environment. Such microbial symbionts therefore likely contribute to local adaptation [1]. The main question underlying the laboratory study by Guilhot et al. [2] was: How much do particular gut bacteria affect the organismal phenotype, in terms of life history and larval foraging traits, of the fruit fly Drosophila melanogaster, a common laboratory model species in biology?
To investigate the above question, the authors isolated 4 taxa of bacteria from the gut of a (randomly picked) Drosophila melanogaster lab strain, and subsequently let Drosophila melanogaster eggs and larvae (stemming from their own, different lab strain) develop both in the typical artificial laboratory medium as well as in grapes, a natural “new” habitat for Drosophila larvae, inoculated with theses bacteria, singly and in combination, also including a bacteria-free control. By investigating various relevant developmental and size traits, the authors found that adding particularly Enterobacteria had some visible effects on several traits, both upward (indicting improvement) and downward (being detrimental) (with three other types of bacteria showing only minor or even no effects). In general, the grape medium reduced performance relative to the standard lab medium. Strongest interactive effects occurred for development time and body size, together making up growth plasticity [3], with lesser such effects on some related behavioral (feeding) traits (Figs. 2,3).
The study premise is interesting, its general objectives are clearly laid out, and the practical work was conducted correctly as far as I can evaluate. The study remains largely descriptive in that no particular a priori hypotheses or predictions in relation to the specific bacteria isolated were formulated, not least because the bacteria were necessarily somewhat arbitrarily chosen and there were apparently no prior studies from which to derive concrete predictions. Overall, the results of this study should be of interest to the community of evolutionary ecologists, especially those working on nutritional and microbiome effects on animal life histories. I consider this work to be primarily ecological, with limited evolutionary content (e.g. no genetics) though some evolutionary implications, as mentioned in the paper’s Conclusions. So this paper would best fit in a microbial or physiological ecology outlet/journal.
The inclusion of a natural medium (grapes) must be commended because this permits inferences and conclusions for at least one natural environment, whereas inferences drawn from laboratory studies in the artificial medium that most Drosophila researchers seem to use are typically limited. Unsurprisingly perhaps, the study showed that Drosophila melanogaster fared generally better in the artificial than the chosen natural medium (grape). Crucially, however, the bacterial symbionts modified both media differentially. Although common bacterial taxa were chosen, the particular bacteria isolated and used remain arbitrary, as there are many. I note that the main and strongest interactive effects between medium and bacterial type are apparent for the Enterobacteria, and they probably also strongly, if not exclusively, mediate the overall effect of the bacterial mixture.
While these specific data are novel, they are not very surprising. If we grow animals in different environments we can expect some detectable effects of these environments, including the bacterial (microbiome) environment, on the hosts life history. The standard and predicted [4] life history response of Drosophila melanogaster (but not all insects [3]) facing stressful nutritional environments, as apparently created by the Enterobacteria, is to extend development but come out smaller in the end. This is what happened here for the laboratory medium ([2]: Fig. 5). The biological interpretation is that individuals have more trouble ingesting and/or digesting the nutrients available (thus prolonging their foraging period and development), yet cannot convert the nutrients effectively into body size increments (hence emerging smaller). This is what the authors here refer to as developmental plasticity, which is ultimately nutritionally mediated. However, interestingly, a signal in the opposite direction was indicated for the bacterial mixture in the grape medium (flies emerging larger after accelerated development: Fig. 5), suggesting some positive effects on growth rate of the natural medium, perhaps related to grapes being a limited resource that needs to be escaped quickly [3]? The reversal of sexual size dimorphism across bacterial treatments in the grape environment detectable in Fig. 4 is interesting, too, though I don’t understand why this happens, and this is not discussed.
In general, more encompassing and increased questions in this context to be researched in the future could be: 1) are these effects predictable (not (yet) at this point, or so it seems); and 2) how strong are these environmental bacterial effects relative to other, more standard effects (e.g. relative to genetic variation, population variation, etc., or relative to other types of environmental effects like, say, temperature)? (3) It could further be asked why not natural but laboratory populations of Drosophila were used for this experiment, if the aim was to draw inferences for the wild situation. (4) Although Genotype x Environment effects are invoked in the Discussion, they were not tested here, lacking genetically different Drosophila families or populations. From an evolutionary standpoint, I consider this the greatest weakness of the study. I was also not too thrilled by the particular statistical analyses employed, though this ultimately does not negate the results. Nevertheless, this work is a good start in this huge field investigating the microbiome. In conclusion, I can recommend this paper after review by PCI Evol Biol.
References
[1] Kawecki, T. J. and Ebert, D. (2004) Conceptual issues in local adaptation. Ecology Letters 7: 1225-1241. doi: 10.1111/j.1461-0248.2004.00684.x
[2] Guilhot, R., Rombaut, A., Xuéreb, A., Howell, K. and Fellous, S. (2019). Environmental specificity in Drosophila-bacteria symbiosis affects host developmental plasticity. BioRxiv, 717702, v3 peer-reviewed and recommended by PCI Evolutionary Biology. doi: 10.1101/717702
[3] Blanckenhorn, W.U. (1999) Different growth responses to temperature and resource limitation in three fly species with similar life histories. Evolutionary Ecology 13: 395-409. doi: 10.1023/A:1006741222586
[4] Stearns, S. C. and Koella, J. (1986) The evolution of phenotypic plasticity in life history traits: predictions of reaction norms for age and size at maturity. Evolution 40: 893-914. doi: 10.1111/j.1558-5646.1986.tb00560.x